1-DAV-202 Data Management 2023/24
Previously 2-INF-185 Data Source Integration
Integrácia dátových zdrojov 2018/19
Website for 2018/19
| 2019-02-21 | (BB) Introduction to Perl (basics, input processing) Lecture 1, Homework 1 |
| 2019-02-28 | (BB) Command-line tools, Perl one-liners Lecture 2, Homework 2 |
| 2019-03-07 | (BB) Job scheduling and make Lecture 3, Homework 3 |
| 2019-03-14 | (BB) Python and SQL for beginners Lecture 4, Homework 4 |
| 2019-03-21 | (VB) Python, web crawling, HTML parsing, sqlite3 Lecture 5 inf, Homework 5 inf |
| (BB) Bioinformatics 1 (genome assembly) Lecture 5 bin, Homework 5 bin | |
| 2019-03-28 | (VB) Text data processing, flask Lecture 6 inf, Homework 6 inf |
| (BB) Bioinformatics 2 (gene finding, RNA-seq) Lecture 6 bin, Homework 6 bin | |
| 2019-04-04 | (VB) Data visualization in JavaScript Lecture 7 inf, Homework inf |
| (BB) Bioinformatics 3 (polymorphisms) Lecture 7 bin, Homework 7 bin | |
| 2019-04-11 | (BB) R, part 1 Lecture 8, Homework 8 |
| 2019-04-18 | Easter (project proposals due Wednesday April 17) |
| 2019-04-25 | (BB) no lecture |
| 2019-05-02 | (BB) R, part 2 Lecture 9, Homework 9 |
| 2019-05-09 | (VB) Cloud computing Lecture 10, Homework 10 |
| 2019-05-16 | no lecture |
Contents
- 1 Kontakt
- 2 Úvod
- 3 Pravidlá
- 4 L01
- 4.1 Why Perl
- 4.2 Hello world
- 4.3 The first input file for today: sequence repeats
- 4.4 A sample Perl program
- 4.5 The second input file for today: DNA sequencing reads (fastq)
- 4.6 Variables, types
- 4.7 Strings, regular expressions
- 4.8 Conditionals, loops
- 4.9 Input, output
- 4.10 Sources of Perl-related information
- 5 HW01
- 6 L02
- 6.1 Efficient use of command line
- 6.2 Redirecting and pipes
- 6.3 Text file manipulation
- 6.3.1 Commands echo and cat (creating and printing files)
- 6.3.2 Commands head and tail (looking at start and end of files)
- 6.3.3 Commands wc, ls -lh, od (exploring file stats and details)
- 6.3.4 Command grep (getting lines matching a regular expression)
- 6.3.5 Commands sort, uniq
- 6.3.6 Commands diff, comm (comparing files)
- 6.3.7 Commands cut, paste, join (working with columns)
- 6.3.8 Commands split, csplit (splitting files to parts)
- 6.4 Programs sed and awk
- 6.5 Perl one-liners
- 7 HW02
- 8 L03
- 9 HW03
- 10 L04
- 11 HW04
- 12 L05inf
- 13 HW05inf
- 14 L05bin
- 15 HW05bin
- 16 L06inf
- 17 HW06inf
- 18 L06bin
- 19 HW06bin
- 20 L07inf
- 21 HW07inf
- 22 L07bin
- 23 HW07bin
- 24 L08
- 25 HW08
- 26 L09
- 27 HW09
- 28 L10
- 29 HW10
Kontakt
Vyučujúci
- doc. Mgr. Broňa Brejová, PhD. miestnosť M-163
- Mgr. Tomáš Vinař, PhD., miestnosť M-163
- Mgr. Vladimír Boža, PhD., miestnosť M-25
- Konzultácie po dohode emailom
Rozvrh
- Štvrtok 15:40-18:00 M-217
Úvod
Cieľová skupina
Tento predmet je určený pre študentov 2. ročníka bakalárskeho študijného programu Bioinformatika a pre študentov bakalárskeho a magisterského študijného programu Informatika, obzvlášť ak plánujú na magisterskom štúdiu absolvovať štátnicové zameranie Bioinformatika a strojové učenie. Radi privítame aj študentov iných zameraní a študijných programov, pokiaľ majú požadované (neformálne) prerekvizity.
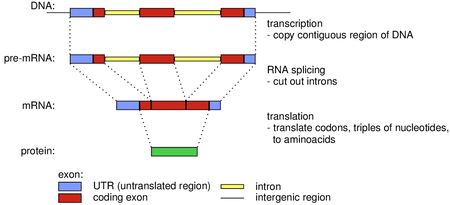
Predpokladáme, že študenti na tomto predmete už vedia programovať v niektorom programovacom jazyku a neboja sa učiť podľa potreby nové jazyky. Takisto predpokladáme základnú znalosť práce v Linuxe vrátane spúšťania príkazov na príkazovom riadku (mali by ste poznať aspoň základné príkazy na prácu so súbormi a adresármi ako cd, mkdir, cp, mv, rm, chmod a pod.). Hoci väčšina technológií preberaných na tomto predmete sa dá použiť na spracovanie dát z mnohých oblastí, budeme ich často ilustrovať na príkladoch z oblasti bioinformatiky. Pokúsime sa vysvetliť potrebné pojmy, ale bolo by dobre, ak by ste sa orientovali v základných pojmoch molekulárnej biológie, ako sú DNA, RNA, proteín, gén, genóm, evolúcia, fylogenetický strom a pod. Študentom zamerania Bioinformatika a strojové učenie odporúčame absolvovať najskôr Metódy v bioinformatike, až potom tento predmet.
Ak sa chcete doučiť základy používania príkazového riadku, skúste napr. tento tutoriál: http://korflab.ucdavis.edu/bootcamp.html
Cieľ predmetu
Počas štúdia sa naučíte mnohé zaujímave algoritmy, modely a metódy, ktoré sa dajú použiť na spracovanie dát v bioinformatike alebo iných oblastiach. Ak však počas štúdia alebo aj neskôr v zamestnaní budete chcieť tieto metódy použiť na reálne dáta, zistíte, že väčšinou treba vynaložiť značné úsilie na samotné získanie dát, ich predspracovanie do vhodného tvaru, testovanie a porovnávanie rôznych metód alebo ich nastavení a získavanie finálnych výsledkov v tvare prehľadných tabuliek a grafov. Často je potrebné tieto činnosti veľakrát opakovať pre rôzne vstupy, rôzne nastavenia a podobne. Obzvlášť v bioinformatike je možné si nájsť zamestnanie, kde vašou hlavnou náplňou bude spracovanie dát s použitím už hotových nástrojov, prípadne doplnených menšími vlastnými programami. Na tomto predmete si ukážeme niektoré programovacie jazyky, postupy a technológie vhodné na tieto činnosti.
Základné princípy
Odporúčame nasledujúci článok s dobrými radami k výpočtovým experimentom:
- Noble WS. A quick guide to organizing computational biology projects. PLoS Comput Biol. 2009 Jul 31;5(7):e1000424.
Niektoré dôležité zásady:
- Citát z článku Noble 2009: "Everything you do, you will probably have to do over again."
- Dobre zdokumentujte všetky kroky experimentu (čo ste robili, prečo ste to robili, čo vám vyšlo)
- Ani vy sami si o pár mesiacov tieto detaily nebudete pamätať
- Snažte sa udržiavať logickú štruktúru adresárov a súborov
- Ak však máte veľa experimentov, môže byť dostačujúce označiť ich dátumom, nevymýšľať stále nové dlhé mená
- Snažte sa vyhýbať manuálnym úpravám medzivýsledkov, ktoré znemožňujú jednoduché zopakovanie experimentu
- Snažte sa detegovať chyby v dátach
- Skripty by mali skončiť s chybovou hláškou, keď niečo nejde ako by malo
- V skriptoch čo najviac kontrolujte, že vstupné dáta zodpovedajú vašim predstavám (správny formát, rozumný rozsah hodnôt atď.)
- Ak v skripte voláte iný program, kontrolujte jeho exit code
- Tiež čo najčastejšie kontrolujte medzivýsledky výpočtu (ručným prezeraním, výpočtom rôznych štatistík a pod.), aby ste odhalili prípadné chyby v dátach alebo vo vašom kóde
Pravidlá
Známkovanie
- Domáce úlohy: 55%
- Návrh projektu: 5%
- Projekt: 40%
Stupnica:
- A: 90 a viac, B:80...89, C: 70...79, D: 60...69, E: 50...59, FX: menej ako 50%
Formát predmetu
- Každý týždeň 3 vyučovacie hodiny, z toho cca prvá je prednáška a ďalšie dve cvičenia. Na cvičeniach samostatne riešite príklady, ktoré doma dokončíte ako domácu úlohu.
- Niektoré týždne bude zvlášť úloha pre študentov bakalárskeho programu Bioinformatika a zvlášť pre ostatných. Ak by ste chceli riešiť iné zadanie, než je pre vás určené, musíte získať vopred súhlas vyučujúcich.
- Cez skúškové obdobie budete odovzdávať projekt. Po odovzdaní projektov sa bude konať ešte diskusia o projekte s vyučujúcimi, ktorá môže ovplyvniť vaše body z projektu.
- Budete mať konto na Linuxovom serveri určenom pre tento predmet. Toto konto používajte len na účely tohto predmetu a snažte sa server príliš svojou aktivitou nepreťažiť, aby slúžil všetkým študentom. Akékoľvek pokusy úmyselne narušiť chod servera budú považované za vážne porušenie pravidiel predmetu.
Domáce úlohy
- Termín DÚ týkajúcej sa aktuálnej prednášky je vždy do 9:00 v deň nasledujúcej prednášky (t.j. väčšinou o necelý týždeň od zadania).
- Domácu úlohu odporúčame začať robiť na cvičení, kde vám môžeme prípadne poradiť. Ak máte otázky neskôr, pýtajte sa vyučujúcich emailom.
- Domácu úlohu môžete robiť na ľubovoľnom počítači, pokiaľ možno pod Linuxom. Odovzdaný kód alebo príkazy by však mali byť spustiteľné na serveri pre tento predmet, nepoužívajte teda špeciálny softvér alebo nastavenia vášho počítača.
- Domáca úloha sa odovzdáva nakopírovaním požadovaných súborov do požadovaného adresára na serveri. Konkrétne požiadavky budú spresnené v zadaní.
- Ak sú mená súborov špecifikované v zadaní, dodržujte ich. Ak ich vymýšľate sami, nazvite ich rozumne. V prípade potreby si spravte aj podadresáre, napr. na jednotlivé príklady.
- Dbajte na prehľadnosť odovzdaného zdrojového kódu (odsadzovanie, rozumné názvy premenných, podľa potreby komentáre)
Protokoly
- Väčšinou bude požadovanou súčasťou úlohy textový dokument nazvaný protokol.
- Protokol môže byť vo formáte .txt alebo .pdf a jeho meno má byť protocol.pdf alebo protocol.txt (nakopírujte ho do odovzdaného adresára)
- Protokol môže byť po slovensky alebo po anglicky.
- V prípade použitia txt formátu a diakritiky ju kódujte v UTF8, ale pre jednoduchosť môžete protokoly písať aj bez diakritiky. Ak je protocol v pdf formáte, mali by sa v ňom dať selektovať texty.
- Vo väčšine úloh dostanete kostru protokolu, dodržujte ju.
Hlavička protokolu, vyhodnotenie
- Na vrchu protokolu uveďte meno, číslo domácej úluhy a vaše vyhodnotenie toho, ako sa vám úlohu podarilo vyriešiť. Vyhodnotenie je prehľadný zoznam všetkých príkladov zo zadania, ktoré ste aspoň začali riešiť a kódov označujúcich ich stupeň dokončenia:
- kód HOTOVO uveďte, ak si myslíte, že tento príklad máte úplne a správne vyriešený
- kód ČASŤ uveďte, ak ste nevyriešili príklad celý a do poznámky za kód stručne uveďte, čo máte hotové a čo nie, prípadne ktorými časťami si nie ste istí.
- kód MOŽNO uveďte, ak príklad máte celý, ale nie ste si istí, či správne. Opäť v poznámke uveďte, čím si nie ste istí.
- kód NIČ uveďte, ak ste príklad ani nezačali riešiť
- Vaše vyhodnotenie je pre nás pomôckou pri bodovaní. Príklady označené HOTOVO budeme kontrolovať námatkovo, k príkladom označeným MOŽNO sa vám pokúsime dať nejakú spätnú väzbu, takisto aj k príkladom označeným ČASŤ, kde v poznámke vyjadríte, že ste mali nejaké problémy.
- Pri vyhodnotení sa pokúste čo najlepšie posúdiť správnosť vašich riešení, pričom kvalita vášho seba-hodnotenia môže vplývať na celkový počet bodov.
Obsah protokolu
- Ak nie je v zadaní určené inak, protokol by mal obsahovať nasledovné údaje:
- Zoznam odovzdaných súborov: o každom súbore uveďte jeho význam a či ste ho vyrobili ručne, získali z externých zdrojov alebo vypočítali nejakým programom. Ak máte väčšie množstvo súborov so systematickým pomenovaním, stačí vysvetliť schému názvov všeobecne. Súbory, ktorých mená sú špecifikované v zadaní, nemusíte v zozname uvádzať.
- Postupnosť všetkých spustených príkazov, prípadne iných krokov, ktorými ste dospeli k získaným výsledkom. Tu uvádzajte príkazy na spracovanie dát a spúšťanie vašich či iných programov. Netreba uvádzať príkazy súvisiace so samotným programovaním (spúšťanie editora, nastavenie práv na spustenie a pod.), s kopírovaním úlohy na server a pod. Uveďte aj stručné komentáre, čo bolo účelom určitého príkazu alebo skupiny príkazov.
- Zoznam zdrojov: webstránky a pod., ktoré ste pri riešení úlohy použili. Nemusíte uvádzať webstránku predmetu a zdroje odporučené priamo v zadaní.
Celkovo by protokol mal umožniť čitateľovi zorientovať sa vo vašich súboroch a tiež v prípade záujmu vykonať rovnaké výpočty, akými ste dospeli vy k výsledku. Nemusíte písať slohy, stačia zrozumiteľné a prehľadné heslovité poznámky.
Projekty
Cieľom projektu je vyskúšať si naučené zručnosti na konkrétnom projekte spracovania dát. Vašou úlohou je zohnať si dáta, tieto dáta analyzovať niektorými technikami z prednášok, prípadne aj inými technológiami a získané výsledky zobraziť v prehľadných grafoch a tabuľkách. Ideálne je, ak sa vám podarí prísť k zaujímavým alebo užitočným záverom, ale hodnotiť budeme hlavne voľbu vhodného postupu a jeho technickú náročnosť. Rozsah samotného programovania alebo analýzy dát by mal zodpovedať zhruba trom domácim úlohám, ale celkovo bude projekt náročnejší, lebo na rozdiel od úloh nemáte postup a dáta vopred určené, ale musíte si ich vymyslieť sami a nie vždy sa prvý nápad ukáže ako správny. V projekte môžete využiť aj existujúce nástroje a knižnice, ale pokiaľ možno používajte nástroje spúšťané na príkazovom riadku.
Zhruba v dvoch tretinách semestra budete odovzdávať návrh projektu (formát txt alebo pdf, rozsah 0.5-1 strana). V tomto návrhu uveďte, aké dáta budete spracovávať, ako ich zoženiete, čo je cieľom analýzy a aké technológie plánujete použiť. Ciele a technológie môžete počas práce na projekte mierne pozmeniť podľa okolností, mali by ste však mať počiatočnú predstavu. K návrhu vám dáme spätnú väzbu, pričom v niektorých prípadoch môže byť potrebné tému mierne alebo úplne zmeniť. Za načas odovzdaný vhodný návrh projektu získate 5% z celkovej známky. Návrh odporúčame pred odovzdaním konzultovať s vyučujúcimi.
Cez skúškové obdobie bude určený termín odovzdania projektu. Podobne ako pri domácich úlohách odovzdávajte adresár s požadovanými súbormi:
- Vaše programy a súbory s dátami (veľmi veľké dátové súbory vynechajte)
- Protokol podobne ako pri domácich úlohách
- formát txt alebo pdf, stručné heslovité poznámky
- obsahuje zoznam súborov, podrobný postup pri analýze dát (spustené príkazy), ako aj použité zdroje (dáta, programy, dokumentácia a iná literatúra atď)
- Správu k projektu vo formáte pdf. Na rozdiel od menej formálneho protokolu by správu mal tvoriť súvislý text v odbornom štýle, podobne ako napr. záverečné práce. Môžete písať po slovensky alebo po anglicky, ale pokiaľ možno gramaticky správne. Správa by mala mať tieto časti:
- úvod, v ktorom vysvetlíte ciele projektu, prípadne potrebné poznatky zo skúmanej oblasti a aké dáta ste mali k dispozícii
- stručný popis metód, v ktorom neuvádzajte detailne jednotlivé kroky, skôr prehľad použitého prístupu a jeho zdôvodnenie
- výsledky analýzy (tabuľky, grafy a pod.) a popis týchto výsledkov, prípadne aké závery sa z nich dajú spraviť (nezabudnite vysvetliť, čo znamenajú údaje v tabuľkách, osi grafov a pod.). Okrem finálnych výsledkov analýzy uveďte aj čiastkové výsledky, ktorými ste sa snažili overovať, že pôvodné dáta a jednotlivé časti vášho postupu sa správajú rozumne.
- diskusiu, v ktorej uvediete, ktoré časti projektu boli náročné a na aké problémy ste narazili, kde sa vám naopak podarilo nájsť spôsob, ako problém vyriešiť jednoducho, ktoré časti projektu by ste spätne odporúčali robiť iným než vašim postupom, čo ste sa na projekte naučili a podobne
Projekty môžete robiť aj vo dvojici, vtedy však vyžadujeme rozsiahlejší projekt a každý člen by mal byť primárne zodpovedný za určitú časť projektu, čo uveďte aj v správe. Dvojice odovzdávajú jednu správu, ale po odovzdaní projektu majú stretnutie s vyučujúcimi individuálne.
Ako nájsť tému projektu:
- Môžete spracovať nejaké dáta, ktoré potrebujete do bakalárskej alebo diplomovej práce, prípadne aj dáta, ktoré potrebujte na iný predmet (v tom prípade uveďte v správe, o aký predmet ide a takisto upovedomte aj druhého vyučujúceho, že ste použili spracovanie dát ako projekt pre tento predmet). Obzvlášť pre BIN študentov môže byť tento predmet vhodnou príležitosťou nájsť si tému bakalárskej práce a začať na nej pracovať.
- Môžete skúsiť zopakovať analýzu spravenú v nejakom vedeckom článku a overiť, že dostanete tie isté výsledky. Vhodné je tiež skúsiť analýzu aj mierne obmeniť (spustiť na iné dáta, zmeniť nejaké nastavenia, zostaviť aj iný typ grafu a pod.)
- Môžete skúsiť nájsť niekoho, kto má dáta, ktoré by potreboval spracovať, ale nevie ako na to (môže ísť o biológov, vedcov z iných oblastí, ale aj neziskové organizácie a pod.) V prípade, že takýmto spôsobom kontaktujete tretie osoby, bolo by vhodné pracovať na projekte obzvlášť zodpovedne, aby ste nerobili zlé meno našej fakulte.
- V projekte môžete porovnávať niekoľko programov na tú istú úlohu z hľadiska ich rýchlosti či presnosti výsledkov. Obsahom projektu bude príprava dát, na ktorých budete programy bežať, samotné spúšťanie (vhodne zoskriptované) ako aj vyhodnotenie výsledkov.
- A samozrejme môžete niekde na internete vyhrabať zaujímavé dáta a snažiť sa z nich niečo vydolovať.
Opisovanie
- Máte povolené sa so spolužiakmi a ďalšími osobami rozprávať o domácich úlohách resp. projektoch a stratégiách na ich riešenie. Kód, získané výsledky aj text, ktorý odovzdáte, musí však byť vaša samostatná práca. Je zakázané ukazovať svoj kód alebo texty spolužiakom.
- Pri riešení domácej úlohy a projektu očakávame, že budete využívať internetové zdroje, najmä rôzne manuály a diskusné fóra k preberaným technológiám. Nesnažte sa však nájsť hotové riešenia zadaných úloh. Všetky použité zdroje uveďte v domácich úlohách a projektoch.
- Ak nájdeme prípady opisovania alebo nepovolených pomôcok, všetci zúčastnení študenti získajú za príslušnú domácu úlohu, projekt a pod. nula bodov (t.j. aj tí, ktorí dali spolužiakom odpísať) a prípad ďalej podstúpime na riešenie disciplinárnej komisii fakulty.
Zverejňovanie
Zadania a materiály k predmetu sú voľne prístupné na tejto stránke. Prosím vás ale, aby ste nezverejňovali ani inak nešírili vaše riešenia domácich úloh, ak nie je v zadaní povedané inak. Vaše projekty môžete zverejniť, pokiaľ to nie je v rozpore s vašou dohodou so zadávateľom projektu a poskytovateľom dát.
L01
This lecture will be an introduction to the Perl scripting language. We will quickly go through some language features, please read the rest of text as work on #HW01.
Why Perl
- From Wikipedia: It has been nicknamed "the Swiss Army chainsaw of scripting languages" because of its flexibility and power, and possibly also because of its "ugliness".
Oficial slogans:
- There's more than one way to do it
- Easy things should be easy and hard things should be possible
Advantages
- Good capabilities for processing text files, regular expressions, running external programs etc.
- Closer to common programming language than shell scripts
- Perl one-liners on the command line can replace many other tools such as sed and awk
- Many existing libraries
Disadvantages
- Quirky syntax
- It is easy to write very unreadable programs (sometimes joking called write-only language)
- Quite slow and uses a lot of memory. If possible do no read entire input to memory, process line by line
Warning: we will use Perl 5, Perl 6 is quite a different language
Hello world
It is possible to run the code directly from a command line (more later):
perl -e'print "Hello world\n"'
This is equivalent to the following code stored in a file:
#! /usr/bin/perl -w use strict; print "Hello world!\n";
- First line is a path to the interpreter
- Swith -w switches warnings on, e.g. if we manipulate with an undefined value (equivalen to "use warnings;")
- Second line use strict will switch on a more strict syntax checks, e.g. all variables must be defined
- Use of -w and use strict is strongly recommended
- Store the program in a file, e.g. hello.pl
- Make it executable (chmod a+x hello.pl)
- Run it with command ./hello.pl
- Also possible to run as perl hello.pl (e.g. if we don't have the path to the interpreter in the file or the executable bit set)
The first input file for today: sequence repeats
- In genomes some sequences occur in many copies (often not exactly equal, only similar)
- We have downloaded a table containing such sequence repeats on chromosome 2L of the fruitfly Drosophila melanogaster
- It was done as follows: on webpage http://genome.ucsc.edu/ we select drosophila genome, then in main menu select Tools, Table browser, select group: variation and repeats, track: ReapatMasker, region: position chr2L, output format: all fields from the selected table a output file: repeats.txt
- Each line of the file contains data about one repeat in the selected chromosome. The first line contains column names. Columns are tab-separated. Here are the first two lines:
#bin swScore milliDiv milliDel milliIns genoName genoStart genoEnd genoLeft strand repName repClass repFamily repStart repEnd repLeft id 585 778 167 7 20 chr2L 1 154 -23513558 + HETRP_DM Satellite Satellite 1519 1669 -203 1
- The file can be found at our server under filename /tasks/hw01/repeats.txt (17185 lines)
- A small randomly selected subset of the table rows is in file /tasks/hw01/repeats-small.txt (159 lines)
A sample Perl program
For each type of repeat (column 11 of the file when counting from 0) we want to compute the number of repeats of this type
#!/usr/bin/perl -w
use strict;
#associative array (hash), with repeat type as key
my %count;
while(my $line = <STDIN>) { # read every line on input
chomp $line; # delete end of line, if any
if($line =~ /^#/) { # skip commented lines
next; # similar to "continue" in C, move to next iteration
}
# split the input line to columns on every tab, store them in an array
my @columns = split "\t", $line;
# check input - should have at least 17 columns
die "Bad input '$line'" unless @columns >= 17;
my $type = $columns[11];
# increase counter for this type
$count{$type}++;
}
# write out results, types sorted alphabetically
foreach my $type (sort keys %count) {
print $type, " ", $count{$type}, "\n";
}
This program does the same thing as the following one-liner (more on one-liners in two weeks)
perl -F'"\t"' -lane 'next if /^#/; die unless @F>=17; $count{$F[11]}++; END { foreach (sort keys %count) { print "$_ $count{$_}" }}' filename
The second input file for today: DNA sequencing reads (fastq)
- DNA sequencing machines can read only short pieces of DNA called reads
- Reads are usually stored in fastq format
- Files can be very large (gigabytes or more), but we will use only a small sample from bacteria Staphylococcus aureus, source [1]
- Each read is on 4 lines:
- line 1: ID of the read and other description, line starts with @
- line 2: DNA sequence, A,C,G,T are bases (nucleotides) of DNA, N means unknown base
- line 3: +
- line 4: quality string, which is the string of the same length as DNA in line 2. Each character represents quality of one base in DNA. If p is the probability that this base is wrong, the quality string will contain character with ASCII value 33+(-10 log p), where log is decimal logarithm. This means that higher ASCII means base of higher quality. Character ! (ASCII 33) means probability 1 of error, character $ (ASCII 36) means 50% error, character + (ASCII 43) is 10% error, character 5 (ASCII 53) is 1% error.
- Note that some sequencing platforms represent qualities differently (see article linked above)
- Our file has all reads of equal length (this is not always the case)
- Technically, a single read and its quality can be split into multiple lines, but this is rarely done, and we will assume that each read takes 4 lines as described above
The first 4 reads from file /tasks/hw01/reads-small.fastq
@SRR022868.1845/1 AAATTTAGGAAAAGATGATTTAGCAACATTTAGCCTTAATGAAAGACCAGATTCTGTTGCCATGTTTGAATGCCTTAAACCAGTAGCAGAATCAGTATAAA + IICIIIIIIIIIID%IIII8>I8III1II,II)I+III*II<II,E;-HI>+I0IB99I%%2GI*=?5*&1>'$0;%'+%%+;#'$&'%%$-+*$--*+(% @SRR022868.1846/1 TAGCGTTGTAAAATAAATTTCTAGAATGGAAGTGATGATATTGAAATACACTCAGATCCTGAATGAAAGATTTATTAAAGTTAAGACGAGAGTCTCATTAT + 4CIIIIIIII52I)IIIII0I16IIIII2IIII;IIAII&I6AI+*+&G5&G.@8/6&%&,03:*.$479.91(9--$,*&/3"$#&*'+#&##&$(&+&+
Variables, types
Scalar variables
- Scalar variables start with $, they can hold undefined value (undef), string, number, reference etc.
- Perl converts automatically between strings and numbers
perl -e'print((1 . "2")+1, "\n")'
13
perl -e'print(("a" . "2")+1, "\n")'
1
perl -we'print(("a" . "2")+1, "\n")'
Argument "a2" isn't numeric in addition (+) at -e line 1.
1
- If we switch on strict parsing, each variable needs to be defined by my, several variables created and initialized as follows: my ($a,$b) = (0,1);
- Usual set of C-style operators, power is **, string concatenation .
- Numbers compared by <, <=, ==, != etc., strings by lt, le, eq, ne, gt, ge,
- Comparison operator $a cmp $b for strings, $a <=> $b for numbers: returns -1 if $a<$b, 0 if they are equal, +1 if $a>$b
Arrays
- Names start with @, e.g. @a
- Access to element 0 in array: $a[0]
- Starts with $, because the expression as a whole is a scalar value
- Length of array scalar(@a). In scalar context, @a is the same thing.
- e.g. for(my $i=0; $i<@a; $i++) { ... }
- If using non-existent indexes, they will be created, initialized to undef (++, += treat undef as 0)
- Stack/vector using functions push and pop: push @a, (1,2,3); $x = pop @a;
- Analogicaly shift and unshift on the left end of the array (slower)
- Sorting
- @a = sort @a; (sorts alphabetically)
- @a = sort {$a <=> $b} @a; (sort numerically)
- { } can contain arbitrary comparison function, $a and $b are the two compared elements
- Array concatenation @c = (@a,@b);
- Swap values of two variables: ($x,$y) = ($y,$x);
- Iterate through values of an array (values can be changed):
perl -e'my @a = (1,2,3); foreach my $val (@a) { $val++; } print join(" ", @a), "\n";'
2 3 4
Associative array (hashes)
- Names start with %, e.g. %b
- Access element with name "X": $b{"X"}
- Write out all elements of associative array %b
foreach my $key (keys %b) {
print $key, " ", $b{$key}, "\n";
}
- Initialization with constant: %b = ("key1"=>"value1", "key2"=>"value2");
- instead of => you can also use ,
- test for existence of a key: if(exists $a{"x"}) {...}
Multidimensional arrays, fun with pointers
- Pointer to a variable: \$a, \@a, \%a
- Pointer to an anonymous array: [1,2,3], pointer to an anonymous hash: {"kluc1"=>"hodnota1"}
- Hash of lists:
my %a = ("fruits"=>["apple","banana","orange"], "vegetables"=>["celery","carrot"]}
$x = $a{"fruits"}[1];
push @{$a{"fruits"}}, "kiwi";
my $aref = \%a;
$x = $aref->{"fruits"}[1];
- Module Data::Dumper has function Dumper, which will recursively print complex data structures
Strings, regular expressions
Strings
- Substring: substr($string, $start, $length)
- used also to access individual charaters (use length 1)
- If we omit $length, considers until the end of the string, negative start counted from the end of the string,...
- We can also replace a substring by something else: substr($str, 0, 1) = "aaa" (replaces the first character by "aaa")
- Length of a string: length($str)
- Splitting a string to parts: split reg_expression, $string, $max_number_of_parts
- if " " instead of regular expression, splits at whitespace
- Connecting parts join($separator, @strings)
- Other useful functions: chomp (removes end of line), index (finds a substring), lc, uc (conversion to lowercase/uppercase), reverse (mirror image), sprintf (C-style formatting)
Regular expressions
- more in [2]
$line =~ s/\s+$//; # remove whitespace at the end of the line
$line =~ s/[0-9]+/X/g; # replace each sequence of numbers with character X
#from the name of the fasta sequence (starting with >) create a string until the first space
#(\S means non-whitespace), the result is stored in $1, as specified by ()
if($line =~ /^\>(\S+)/) { $name = $1; }
perl -le'$X="123 4 567"; $X=~s/[0-9]+/X/g; print $X'
X X X
Conditionals, loops
if(expression) { # () and {} cannot be omitted
commands
} elsif(expression) {
commands
} else {
commands
}
command if expression; # here () not necessary
command unless expression;
die "negative value of x: $x" unless $x>=0;
for(my $i=0; $i<100; $i++) {
print $i, "\n";
}
foreach my $i (0..99) {
print $i, "\n";
}
$x=1;
while(1) {
$x *= 2;
last if $x>=100;
}
- Undefined value, number 0 and strings "" and "0" evaluate as false, but I would recommmend always explicitly using logical values in conditional expressions, e.g. if(defined $x), if($x eq ""), if($x==0) etc.
Input, output
- Reading one line from standard input: $line = <STDIN>
- If no more input data available, returns undef
- See also [3]
- Special idiom while(my $line = <STDIN>) equivalent to while (defined(my $line = <STDIN>))
- iterates through all lines of input
- chomp $line removes "\n", if any from the end of the string
- output to stdout through print or printf
- Lecture Perl 2 in IDZ archive (files, command-line arguments, running external programs, defining functions abd modules, Bioperl)
- In package perl-doc man pages:
- man perlintro introduction to Perl
- man perlfunc list of standard functions in Perl
- perldoc -f split describes function split, similarly other functions
- perldoc -q sort shows answers to commonly asked questions (FAQ)
- man perlretut and man perlre regular expressions
- man perl list of other manual pages about Perl
- The same content on the web http://perldoc.perl.org/
- Various web tutorials e.g. this one
- Books
- Bioperl [6] big library for bioinformatics
- Perl for Windows: http://strawberryperl.com/
HW01
See Lecture 1
Files
We have 4 input files for this homework. We recommend creating soft links to your working directory as follows:
ln -s /tasks/hw01/repeats-small.txt . # small version of the repeat file ln -s /tasks/hw01/repeats.txt . # full version of the repeat file ln -s /tasks/hw01/reads-small.fastq . # smaller version of the read file ln -s /tasks/hw01/reads.fastq . # bigger version of the read file
We recommend writing your protocol starting from an outline provided in /tasks/hw01/protocol.txt
Submitting
- Directory /submit/hw01/your_username will be created for you
- Copy required files to this directory, including the protocol named protocol.txt or protocol.pdf
- You can modify these files freely until deadline, but after the deadline of the homework, you will lose access rights to this directory
Task A
- Consider the program for counting repeat types in the lecture 1, save it to file repeat-stat.pl
- Extend it to compute the average length of each type of repeat
- Each row of the input table contains the start and end coordinates of the repeat in columns 7 and 6. The length is simply the difference of these two values.
- Output a table with three columns: type of repeat, the number of occurrences, the average length of the repeat.
- Use printf to print these three items right-justified in columns of sufficient width, print the average length to 1 decimal place.
- If you run your script on the small file, the output should look something like this (exact column widths may differ):
./repeat-stat.pl < repeats-small.txt
DNA 5 377.4
LINE 4 410.2
LTR 13 355.4
Low_complexity 22 47.2
RC 8 236.2
Simple_repeat 106 39.0
- Include in your protocol the output when you run your script on the large file: ./repeat-stat.pl < repeats.txt
- Find out on Wikipedia, what acronyms LINE and LTR stand for. Do their names correspond to their lengths? (Write a short answer in the protocol.)
- Submit only your script, repeat-stat.pl
Task B
- Write a script which reformats FASTQ file to FASTA format, call it fastq2fasta.pl
- fastq file should be on standard input, fasta file written to standard output
- FASTA format is a typical format for storing DNA and protein sequences.
- Each sequence consists of several lines of the file. The first line starts with ">" followed by identifier of the sequence and optionally some further description separated by whitespace
- The sequence itself is on the second line, long sequences are split into multiple lines
- In our case, the name of the sequence will be the ID of the read with @ replaced by > and / replaced by _
- you can try to use tr or s operators (see also lecture)
- For example, the first two reads of reads.fastq are:
@SRR022868.1845/1 AAATTTAGGAAAAGATGATTTAGCAACATTTAGCCTTAATGAAAGACCAGATTCTGTTGCCATGTTTGAATGCCTTAAACCAGTAGCAGAATCAGTATAAA + IICIIIIIIIIIID%IIII8>I8III1II,II)I+III*II<II,E;-HI>+I0IB99I%%2GI*=?5*&1>'$0;%'+%%+;#'$&'%%$-+*$--*+(% @SRR022868.1846/1 TAGCGTTGTAAAATAAATTTCTAGAATGGAAGTGATGATATTGAAATACACTCAGATCCTGAATGAAAGATTTATTAAAGTTAAGACGAGAGTCTCATTAT + 4CIIIIIIII52I)IIIII0I16IIIII2IIII;IIAII&I6AI+*+&G5&G.@8/6&%&,03:*.$479.91(9--$,*&/3"$#&*'+#&##&$(&+&+
- These should be reformatted as follows:
>SRR022868.1845_1 AAATTTAGGAAAAGATGATTTAGCAACATTTAGCCTTAATGAAAGACCAGATTCTGTTGCCATGTTTGAATGCCTTAAACCAGTAGCAGAATCAGTATAAA >SRR022868.1846_1 TAGCGTTGTAAAATAAATTTCTAGAATGGAAGTGATGATATTGAAATACACTCAGATCCTGAATGAAAGATTTATTAAAGTTAAGACGAGAGTCTCATTAT
- Submit files fastq2fasta.pl and reads-small.fasta
- the latter file is created by running ./fastq2fasta.pl < reads-small.fastq > reads-small.fasta
Task C
- Write a script fastq-quality.pl which for each position in a read computes the average quality
- Standard input has fastq file with multiple reads, possibly of different lengths
- As quality we will use ASCII values of characters in the quality string with value 33 subtracted, so the quality is -10 log p
- ASCII value can be computed by function ord
- Positions in reads will be numbered from 0
- Since reads can differ in length, some positions are used in more reads, some in fewer
- For each position from 0 up to the highest position used in some read, print three numbers separated by tabs "\t": the position index, the number of times this position was used in reads, the average quality at that position with 1 decimal place (you can again use printf)
- The last two lines when you run ./fastq-quality.pl < reads-small.fastq should be
99 86 5.5 100 86 8.6
- Run the following command, which runs your script on the larger file and selects every 10th position. Include the output in your protocol. Do you see any trend in quality values with increasing position? (Include a short comment in protocol.)
./fastq-quality.pl < reads.fastq | perl -lane 'print if $F[0]%10==0'
- Submit only fastq-quality.pl
Task D
- Write script fastq-trim.pl that trims low quality bases from the end of each read and filters out short reads
- This script should read a fastq file from standard input and write trimmed fastq file to standard output
- It should also accept two command-line arguments: character Q and integer L
- We have not covered processing command line arguments, but you can use the code snippet below
- Q is the minimum acceptable quality (characters from quality string with ASCII value >= ASCII value of Q are ok)
- L is the minimum acceptable length of a read
- First find the last base in a read which has quality at least Q (if any). All bases after this base will be removed from both the sequence and quality string
- If the resulting read has fewer than L bases, it is omitted from the output
You can check your program by the following tests:
- If you run the following two commands, you should get file tmp identical with input and thus output of diff should be empty
./fastq-trim.pl '!' 101 < reads-small.fastq > tmp # trim at quality ASCII >=33 and length >=101 diff reads-small.fastq tmp # output should be empty (no differences)
- If you run the following two commands, you should see differences in 4 reads, 2 bases trimmed from each
./fastq-trim.pl '"' 1 < reads-small.fastq > tmp # trim at quality ASCII >=34 and length >=1 diff reads-small.fastq tmp # output should be differences in 4 reads
- If you run the following commands, you should get empty output (no reads meet the criteria):
./fastq-trim.pl d 1 < reads-small.fastq # quality ASCII >=100, length >= 1 ./fastq-trim.pl '!' 102 < reads-small.fastq # quality ASCII >=33 and length >=102
Further runs and submitting
- Run ./fastq-trim.pl '(' 95 < reads-small.fastq > reads-small-filtered.fastq # quality ASCII >= 40
- Submit files fastq-trim.pl and reads-small-filtered.fastq
- If you have done task C, run quality statistics on the trimmed version of the bigger file using command below and include the result in the protocol. Comment in the protocol on the differences between statistics on the whole file in part C and D. Are they as you expected?
./fastq-trim.pl 2 50 < reads.fastq | ./fastq-quality.pl | perl -lane 'print if $F[0]%10==0' # quality ASCII >= 50
- Note: you have created tools which can be combined, e.g. you can first trim fastq and then convert it to fasta (no need to submit these files)
Parsing command-line arguments in this task (they will be stored in variables $Q and $L):
#!/usr/bin/perl -w use strict; my $USAGE = " Usage: $0 Q L < input.fastq > output.fastq Trim from the end of each read bases with ASCII quality value less than the given threshold Q. If the length of the read after trimming is less than L, the read will be omitted from output. L is a non-negative integer, Q is a character "; # check that we have exactly 2 command-line arguments die $USAGE unless @ARGV==2; # copy command-line arguments to variables Q and L my ($Q, $L) = @ARGV; # check that $Q is one character and $L looks like a non-negative integer die $USAGE unless length($Q)==1 && $L=~/^[0-9]+$/;
L02
Today: using command-line tools and Perl one-liners.
- We will do simple transformations of text files using command-line tools without writing any scripts or longer programs.
- You will record the commands used in your protocol
- We strongly recommend making a log of commands for data processing also outside of this course
- If you have a log of executed commands, you can easily execute them again by copy and paste
- For this reason any comments are best preceded by #
- If you use some sequence of commands often, you can turn it into a script
Efficient use of command line
Some tips for bash shell:
- use tab key to complete command names, path names etc
- tab completion can be customized [7]
- use up and down keys to walk through history of recently executed commands, then edit and resubmit chosen command
- press ctrl-r to search in the history of executed commands
- at the end of session, history stored in ~/.bash_history
- command history -a appends history to this file right now
- you can then look into the file and copy appropriate commands to your protocol
- various other history tricks, e.g. special variables [8]
- cd - goes to previously visited directory, also see pushd and popd
- ls -lt | head shows 10 most recent files, useful for seeing what you have done last
Instead of bash, you can use more advanced command-line environments, e.g. iPhyton notebook
Redirecting and pipes
# redirect standard output to file command > file # append to file command >> file # redirect standard error command 2>file # redirect file to standard input command < file # do not forget to quote > in other uses, e.g. when searching for string ">" in a file sequences.fasta grep '>' sequences.fasta # (without quotes rewrites sequences.fasta) # other special characters, such as ;, &, |, # etc should be quoted in '' as well # send stdout of command1 to stdin of command2 command1 | command2 # backtick operator executes command, # removes trailing \n from stdout, substitutes to command line # the following commands do the same thing: head -n 2 file head -n `echo 2` file # redirect a string in ' ' to stdin of command head head -n 2 <<< 'line 1 line 2 line 3' # in some commands, file argument can be taken from stdin if denoted as - or stdin or /dev/stdin # the following compares uncompressed version of file1 with file2 zcat file1.gz | diff - file2
Make piped commands fail properly:
set -o pipefail
If set, the return value of a pipeline is the value of the last (rightmost) command to exit with a non-zero status, or zero if all commands in the pipeline exit successfully. This option is disabled by default, pipe then returns exit status of the rightmost command.
Text file manipulation
Commands echo and cat (creating and printing files)
# print text Hello and end of line to stdout echo "Hello" # interpret backslash combinations \n, \t etc: echo -e "first line\nsecond\tline" # concatenate several files to stdout cat file1 file2
Commands head and tail (looking at start and end of files)
# print 10 first lines of file (or stdin) head file some_command | head # print the first 2 lines head -n 2 file # print the last 5 lines tail -n 5 file # print starting from line 100 (line numbering starts at 1) tail -n +100 file # print lines 81..100 head -n 100 file | tail -n 20
Commands wc, ls -lh, od (exploring file stats and details)
# prints three numbers: number of lines (-l), number of words (-w), number of bytes (-c) wc file # prints size of file in human-readable units (K,M,G,T) ls -lh file # od -a prints file or stdout with named characters # allows checking whitespace and special characters echo "hello world!" | od -a # prints: # 0000000 h e l l o sp w o r l d ! nl # 0000015
Command grep (getting lines matching a regular expression)
# -i ignores case (upper case and lowercase letters are the same) grep -i chromosome file # -c counts the number of matching lines in each file grep -c '^[12][0-9]' file1 file2 # other options (there is more, see the manual): # -v print/count not matching lines (inVert) # -n show also line numbers # -B 2 -A 1 print 2 lines before each match and 1 line after match # -E extended regular expressions (allows e.g. |) # -F no regular expressions, set of fixed strings # -f patterns in a file # (good for selecting e.g. only lines matching one of "good" ids)
- docs: grep
Commands sort, uniq
# some useful options of sort: # -g numeric sort # -k which column(s) to use as key # -r reverse (from largest values) # -s stable # -t fields separator # sorting first by column 2 numerically (-k2,2g), in case of ties use column 1 (-k1,1) sort -k2,2g -k1,1 file # uniq outputs one line from each group of consecutive identical lines # uniq -c adds the size of each group as the first column # the following finds all unique lines and sorts them by frequency from the most frequent sort file | uniq -c | sort -gr
Commands diff, comm (comparing files)
diff compares two files, useful for manual checking of differences
- useful options
- -b (ignore whitespace differences)
- -r for comparing whole directories
- -q for fast checking for identity
- -y show differences side-by-side
comm compares two sorted files
- writes 3 columns:
- 1: lines occurring only in the first file
- 2: lines occurring only in the second file
- 3: lines occurring in both files
- some columns can be suppressed with -1, -2, -3
- good for finding set intersections and differences
Commands cut, paste, join (working with columns)
- cut selects only some columns from file (perl/awk more flexible)
- paste puts 2 or more files side by side, separated by tabs or other character
- join is a powerful tool for making joins and left-joins as in databases on specified columns in two files
Commands split, csplit (splitting files to parts)
- split splits into fixed-size pieces (size in lines, bytes etc.)
- csplit splits at occurrence of a pattern (e.g. fasta file into individual sequences)
csplit sequences.fa '/^>/' '{*}'
Programs sed and awk
Both programs process text files line by line, allow to do various transformations
# replace text "Chr1" by "Chromosome 1"
sed 's/Chr1/Chromosome 1/'
# prints first two lines, then quits (like head -n 2)
sed 2q
# print first and second column from a file
awk '{print $1, $2}'
# print the line if difference in first and second column > 10
awk '{ if ($2-$1>10) print }'
# print lines matching pattern
awk '/pattern/ { print }'
# count lines
awk 'END { print NR }'
Perl one-liners
Instead of sed and awk, we will cover Perl one-liners
# -e executes commands
perl -e'print 2+3,"\n"'
perl -e'$x = 2+3; print $x, "\n"';
# -n wraps commands in a loop reading lines from stdin or files listed as arguments
# the following is roughly the same as cat:
perl -ne'print'
# how to use:
perl -ne'print' < input > output
perl -ne'print' input1 input2 > output
# lines are stored in a special variable $_
# this variable is default argument of many functions,
# including print, so print is the same as print $_
# simple grep-like commands:
perl -ne 'print if /pattern/'
# simple regular expression modifications
perl -ne 's/Chr(\d+)/Chromosome $1/; print'
# // and s/// are applied by default to $_
# -l removes end of line from each input line and adds "\n" after each print
# the following adds * at the end of each line
perl -lne'print $_, "*"'
# -a splits line into words separated by whitespace and stores them in array @F
# the next example prints difference in numbers stored in the second and first column
# (e.g. interval size if each line coordinates of one interval)
perl -lane'print $F[1]-$F[0]'
# -F allows to set separator used for splitting (regular expression)
# the next example splits at tabs
perl -F '"\t"' -lane'print $F[1]-$F[0]'
# END { commands } is run at the very end, after we finish reading input
# the following example computes the sum of interval lengths
perl -lane'$sum += $F[1]-$F[0]; END { print $sum; }'
# similarly BEGIN { command } before we start
Other interesting possibilites:
# -i replaces each file with a new transformed version (DANGEROUS!)
# the next example removes empty lines from all .txt files in the current directory
perl -lne 'print if length($_)>0' -i *.txt
# the following example replaces sequence of whitespace by exactly one space
# and removes leading and trailing spaces from lines in all .txt files
perl -lane 'print join(" ", @F)' -i *.txt
# variable $. contains line number. $ARGV name of file or - for stdin
# the following prints filename and line number in front of every line
perl -ne'printf "%s.%d: %s", $ARGV, $., $_' file1 file2
# moving files *.txt to have extension .tsv:
# first print commands
# then execute by hand or replace print with system
# mv -i asks if something is to be rewritten
ls *.txt | perl -lne '$s=$_; $s=~s/\.txt/.tsv/; print("mv -i $_ $s")'
ls *.txt | perl -lne '$s=$_; $s=~s/\.txt/.tsv/; system("mv -i $_ $s")'
HW02
Lecture 1 (Perl 1), Lecture 2 (command-line)
- In this homework, use command-line tools or one-liners in Perl, awk or sed. Do not write any scripts or programs.
- Each task can be split into several stages and intermediate files written to disk, but you can also use pipelines to reduce the number of temporary files.
- Your commands should work also for other input files with the same format (do not try to generalize them too much, but also do not use very specific properties of a particular input, such as the number of lines etc.)
- Include all relevant used commands in your protocol and add a short description of your approach.
- Submit the protocol and required output files.
- Outline of the protocol is in /tasks/hw02/protocol.txt, submit to directory /submit/hw02/yourname
Task A
- /tasks/hw02/names.txt contains data about several people, one per line.
- Each line consists of given name(s), surname and email separated by spaces.
- Each person can have multiple given names (at least 1), but exactly one surname and one email. Email is always of the form username@uniba.sk.
- The task is to generate file passwords.csv which contains a randomly generated password for each of these users
- The output file has columns separated by commas ','
- The first column contains username extracted from email address, the second column surname, the third column all given names and the fourth column the randomly generated password
- Submit file passwords.csv with the result of your commands.
Example line from input:
Pavol Országh Hviezdoslav hviezdoslav32@uniba.sk
Example line from output (password will differ):
hviezdoslav32,Hviezdoslav,Pavol Országh,3T3Pu3un
Hints:
- Passwords can be generated using pwgen (e.g. pwgen -N 10 -1 prints 10 passwords, one per line)
- We also recommend using perl, wc, paste (check option -d in paste)
- In Perl, function pop may be useful for manipulating @F and function join for connecting strings with a separator.
Task B
File:
- /tasks/hw02/saccharomyces_cerevisiae.gff contains annotation of the yeast genome
- Downloaded from http://yeastgenome.org/ on 2016-03-09, in particular from [12].
- It was further processed to omit DNA sequences from the end of file.
- The size of the file is 5.6M.
- For easier work, link the file to your directory by ln -s /tasks/hw02/saccharomyces_cerevisiae.gff yeast.gff
- The file is in GFF3 format [13]
- Lines starting with # are comments, other lines contain tab-separated data about one interval of some chromosome in the yeast genome
- Meaning of the first 5 columns:
- column 0 chromosome name
- column 1 source (can be ignored)
- column 2 type of interval
- column 3 start of interval (1-based coordinates)
- column 4 end of interval (1-based coordinates)
- You can assume that these first 5 columns do not contain whitespace
Task:
- Print for each type of interval (column 2), how many times it occurs in the file.
- Sort from the most common to the least common interval types.
- Hint: commands sort and uniq will be useful. Do not forget to skip comments, for example using grep -v '^#'
- Submit file types.txt with the output formatted as follows:
7058 CDS
6600 mRNA
...
...
1 telomerase_RNA_gene
1 mating_type_region
1 intein_encoding_region
Task C
- Continue processing file from task B.
- For each chromosome, the file contains a line which has in column 2 string chromosome, and the interval is the whole chromosome.
- To file chrosomes.txt, print a tab-separated list of chromosome names and sizes in the same order as in the input
- The last line of chromosomes.txt should list the total size of all chromosomes combined.
- Submit file chromosomes.txt
- Hints:
- The total size can be computed by a perl one-liner.
- Example from the lecture: compute the sum of interval sizes if each line of the file contains start and end of one interval: perl -lane'$sum += $F[1]-$F[0]; END { print $sum; }'
- Grepping for word chromosome does not check if this word is indeed in the second column
- Tab character is written in Perl as "\t".
- Your output should start and end as follows:
chrI 230218 chrII 813184 ... ... chrXVI 948066 chrmt 85779 total 12157105
Task D
Overall goal:
- Proteins from several well-studied yeast species were downloaded from database http://www.uniprot.org/ on 2016-03-09
- We have also downloaded proteins from yeast Yarrowia lipolytica. We will pretend that nothing is known about these proteins (as if they were produced by gene finding program in a newly sequenced genome).
- For each Y.lip. proteins we have found similar proteins from other yeasts
- Now we want to find for each protein in Y.lip. its closest match among all known proteins.
Files:
- /tasks/hw02/known.fa is a fasta file with known proteins from several species
- /tasks/hw02/yarLip.fa is a fasta file with proteins from Y.lip.
- /tasks/hw02/known.blast is the result of finding similar proteins in yarLip.fa versus known.fa by these commands:
formatdb -i known.fa blastall -p blastp -d known.fa -i yarLip.fa -m 9 -e 1e-5 > known.blast
- you can link these files to your directory as follows:
ln -s /tasks/hw02/known.fa . ln -s /tasks/hw02/yarLip.fa . ln -s /tasks/hw02/known.blast .
Step 1:
- Get the first (strongest) match for each query from known.blast.
- This can be done by printing the lines that are not comments but follow a comment line starting with #.
- In a perl one-liner, you can create a state variable which will remember if the previous line was a comment and based on that you decide of you print the current line.
- Instead of using perl, you can play with grep. Option -A 1 prints the matching lines as well as one line ofter each match
- Print only the first two columns separated by tab (name of query, name of target), sort the file by the second column.
- Submit file best.tsv with the result
- File should start as follows:
Q6CBS2 sp|B5BP46|YP52_SCHPO Q6C8R4 sp|B5BP48|YP54_SCHPO Q6CG80 sp|B5BP48|YP54_SCHPO Q6CH56 sp|B5BP48|YP54_SCHPO
Step 2:
- Submit file known.tsv which contains sequence names extracted from known.fa with leading > removed
- This file should be sorted alphabetically.
- File should start as follows:
sp|A0A023PXA5|YA19A_YEAST Putative uncharacterized protein YAL019W-A OS=Saccharomyces cerevisiae (strain ATCC 204508 / S288c) GN=YAL019W-A PE=5 SV=1 sp|A0A023PXB0|YA019_YEAST Putative uncharacterized protein YAR019W-A OS=Saccharomyces cerevisiae (strain ATCC 204508 / S288c) GN=YAR019W-A PE=5 SV=1
Step 3:
- Use command join to join the files best.tsv and known.tsv so that each line of best.tsv is extended with the text describing the corresponding target in known.tsv
- Use option -1 2 to use the second column of best.tsv as a key for joining
- The output of join may look as follows:
sp|B5BP46|YP52_SCHPO Q6CBS2 Putative glutathione S-transferase C1183.02 OS=Schizosaccharomyces pombe (strain 972 / ATCC 24843) GN=SPBC460.02c PE=3 SV=1 sp|B5BP48|YP54_SCHPO Q6C8R4 Putative alpha-ketoglutarate-dependent sulfonate dioxygenase OS=Schizosaccharomyces pombe (strain 972 / ATCC 24843) GN=SPBC460.04c PE=3 SV=1
- Further reformat the output so that query name goes first (e.g. Q6CBS2), followed by target name (e.g. sp|B5BP46|YP52_SCHPO), followed by the rest of the text, but remove all text after OS=
- Sort by query name
- Submit file best.txt with the result
- The output should start as follows:
B5FVA8 tr|Q5A7D5|Q5A7D5_CANAL Lysophospholipase B5FVB0 sp|O74810|UBC1_SCHPO Ubiquitin-conjugating enzyme E2 1 B5FVB1 sp|O13877|RPAB5_SCHPO DNA-directed RNA polymerases I, II, and III subunit RPABC5
Note:
- Not all Y.lip. are necessarily included in your final output (some proteins do not have blast match).
- You can think how to find the list of such proteins, but this is not part of the assignment.
- Files best.txt and best.tsv should have the same number of lines.
L03
Job Scheduling
- Some computing jobs take a lot of time: hours, days, weeks,...
- We do not want to keep a command-line window open the whole time; therefore we run such jobs in the background
- Simple commands to do it in Linux:
- Now we will concentrate on Sun Grid Engine, a complex software for managing many jobs from many users on a cluster from multiple computers
- Basic workflow:
- Submit a job (command) to a queue
- The job waits in the queue until resources (memory, CPUs, etc.) become available on some computer
- The job runs on the computer
- Output of the job is stored in files
- User can monitor the status of the job (waiting, running)
- Complex possibilities for assigning priorities and deadlines to jobs, managing multiple queues etc.
- Ideally all computers in the cluster share the same environment and filesystem
- We have a simple training cluster for this exercise:
- You submit jobs to queue on vyuka
- They will run on computer cpu02
- This cluster is only temporarily available until next Thursday
Submitting a job (qsub)
- qsub -b y -cwd 'command < input > output 2> error'
- quoting around command allows us to include special characters, such as <, > etc. and not to apply it to qsub command itself
- -b y treats command as binary, usually preferable for both binary programs and scripts
- -cwd executes command in the current directory
- -N name allows to set name of the job
- -l resource=value requests some non-default resources
- for example, we can use -l threads=2 to request 2 threads for parallel programs
- Grid engine will not check if you do not use more CPUs or memory than requested, be considerate (and perhaps occasionally watch your jobs by running top at the computer where they execute)
- qsub will create files for stdout and stderr, e.g. s2.o27 and s2.e27 for the job with name s2 and jobid 27
Monitoring and deleting jobs (qstat, qdel)
- qstat displays jobs of the current user
job-ID prior name user state submit/start at queue slots ja-task-ID
-----------------------------------------------------------------------------------------------------------------
28 0.50000 s3 bbrejova r 03/15/2016 22:12:18 main.q@cpu02.compbio.fmph.unib 1
29 0.00000 s3 bbrejova qw 03/15/2016 22:14:08 1
- qstat -u '*' displays jobs of all users
- finished jobs disappear from the list
- qstat -F threads shows how many threads available
queuename qtype resv/used/tot. load_avg arch states
---------------------------------------------------------------------------------
main.q@cpu02.compbio.fmph.unib BIP 0/2/8 0.03 lx26-amd64
hc:threads=0
28 0.75000 s3 bbrejova r 03/15/2016 22:12:18 1
29 0.25000 s3 bbrejova r 03/15/2016 22:14:18 1
- Command qdel allows you to delete a job (waiting or running)
Interactive work on the cluster (qrsh), screen
- qrsh creates a job which is a normal interactive shell running on the cluster
- in this shell you can manually run commands
- when you close the shell, the job finishes
- therefore it is a good idea to run qrsh within screen
- run screen command, this creates a new shell
- within this shell, run qrsh, then whatever commands
- by pressing Ctrl-a d you "detach" the screen, so that both shells (local and qrsh) continue running but you can close your local window
- later by running screen -r you get back to your shells
Running many small jobs
For example, consider tens of thousands of genes, run some computation for each gene
- Have a script which iterates through all and runs them sequentially
- Problems: Does not use parallelism, needs more programming to restart after some interruption
- Submit processing of each gene as a separate job to cluster (submitting done by a script/one-liner)
- Jobs can run in parallel on many different computers
- Problem: Queue gets very long, hard to monitor progress, hard to resubmit only unfinished jobs after some failure.
- Array jobs in qsub (option -t): runs jobs numbered 1,2,3...; number of the job is in an environment variable, used by the script to decide which gene to process
- Queue contains only running sub-jobs plus one line for the remaining part of the array job.
- After failure, you can resubmit only unfinished portion of the interval (e.g. start from job 173).
- Next: using make in which you specify how to process each gene and submit a single make command to the queue
- Make can execute multiple tasks in parallel using several threads on the same computer (qsub array jobs can run tasks on multiple computers)
- It will automatically skip tasks which are already finished
Make
- Make is a system for automatically building programs (running compiler, linker etc)
- In particular, we will use GNU make
- Rules for compilation are written in a Makefile
- Rather complex syntax with many features, we will only cover basics
Rules
- The main part of a Makefile are rules specifying how to generate target files from some source files (prerequisites).
- For example the following rule generates target.txt by concatenating source1.txt a source2.txt:
target.txt : source1.txt source2.txt
cat source1.txt source2.txt > target.txt
- The first line describes target and prerequisites, starts in the first column
- The following lines list commands to execute to create the target
- Each line with a command starts with a tab character
- If we have a directory with this rule in Makefile and files source1.txt and source2.txt, running make target.txt will run the cat command
- However, if target.txt already exists, the command will be run only if one of the prerequisites has more recent modification time than the target
- This allows to restart interrupted computations or rerun necessary parts after modification of some input files
- Makefile automatically chains the rules as necessary:
- if we run make target.txt and some prerequisite does not exist, Makefile checks if it can be created by some other rule and runs that rule first
- In general it first finds all necessary steps and runs them in topological order so that each rules has its prerequisites ready
- Option make -n target will show what commands would be executed to build target (dry run) - good idea before running something potentially dangerous
Pattern rules
- We can specify a general rule for files with a systematic naming scheme. For example, to create a .pdf file from a .tex file, we use pdflatex command:
%.pdf : %.tex
pdflatex $^
- In the first line, % denotes some variable part of the filename, which has to agree in the target and all prerequisites
- In commands, we can use several variables:
- $^ contains name for the prerequisite (source)
- $@ contains the name of the target
- $* contains the string matched by %
Other useful tricks in Makefiles
Variables
- Store some reusable values in variables, then use them several times in the Makefile:
MYPATH := /projects/trees/bin
target : source
$(MYPATH)/script < $^ > $@
Wildcards, creating a list of targets from files in the directory
The following Makefile automatically creates .png version of each .eps file simply by running make:
EPS := $(wildcard *.eps)
EPSPNG := $(patsubst %.eps,%.png,$(EPS))
all: $(EPSPNG)
clean:
rm $(EPSPNG)
%.png : %.eps
convert -density 250 $^ $@
- variable EPS contains names of all files matching *.eps
- variable EPSPNG contains desirable names of png files
- it is created by taking filenames in EPS and changing .eps to .png
- all is a "phony target" which is not really created
- its rule has no commands but all png files are prerequisites, so are done first
- the first target in Makefile (in this case all) is default when no other target is specified on command-line
- clean is also a phony target for deleting generated png files
Useful special built-in target names
Include these lines in your Makefile if desired
.SECONDARY: # prevents deletion of intermediate targets in chained rules .DELETE_ON_ERROR: # delete targets if a rule fails
Parallel make
- running make with option -j 4 will run up to 4 commands in parallel if their dependencies are already finished
- easy parallelization on a single computer
Alternatives to Makefiles
- Bioinformatics often uses "pipelines" - sequences of commands run one after another, e.g. by a script of Makefile
- There are many tools developed for automating computational pipelines, see e.g. this review: Jeremy Leipzig; A review of bioinformatic pipeline frameworks. Brief Bioinform 2016 bbw020.
- For example Snakemake
- Workflows can contain shell commands or Python code
- Big advantage compared to Make: pattern rules may contain multiple variable portions (in make only one % per filename)
- For example, you have several fasta files and several HMMs representing protein families and you wans to run each HMM on each fasta file:
rule HMMER:
input: "{filename}.fasta", "{hmm}.hmm"
output: "{filename}_{hmm}.hmmer"
shell: "hmmsearch --domE 1e-5 --noali --domtblout {output} {input[1]} {input[0]}"
HW03
See also Lecture 3
Motivation: Building Phylogenetic Trees
The task for today will be to build a phylogenetic tree of 9 mammalian species using protein sequences
- A phylogenetic tree is a tree showing evolutionary history of these species. Leaves are target present-day species, internal nodes are their common ancestors.
- Input contains sequences of selected proteins from each species
- Step 1: Identify ortholog groups. Orthologs are proteins from different species that "correspond" to each other. This is done based on sequence similarity and we can use a tool called blast to identify sequence similarities between individual proteins. The result of ortholog group identification will be a set of groups, each group having one sequence from each of the 9 species
- Step 2: For each ortholog group, we need to align proteins and build a phylogenetic tree for this protein using existing methods. We can do this using tools muscle (for alignment) and phyml (for phylogenetic tree inference).
Unaligned sequences (start of protein O60568):
>human MTSSGPGPRFLLLLPLLLPPAASASDRPRGRDPVNPEKLLVITVA... >baboon MTSSRPGLRLLLLLLLLPPAASASDRPRGRDPVNPEKLLVMTVA... >dog MASSGPGLRLLLGLLLLLPPPPATSASDRPRGGDPVNPEKLLVITVA... >elephant MASWGPGARLLLLLLLLLLPPPPATSASDRSRGSDRVNPERLLVITVA... >guineapig MAFGAWLLLLPLLLLPPPPGACASDQPRGSNPVNPEKLLVITVA... >opossum SDKLLVITAA... >pig AMASGPGLRLLLLPLLVLSPPPAASASDRPRGSDPVNPDKLLVITVA... >rabbit MGCDSRKPLLLLPLLPLALVLQPWSARGRASAEEPSSISPDKLLVITVA... >rat MAASVPEPRLLLLLLLLLPPLPPVTSASDRPRGANPVNPDKLLVITVA...
Aligned sequences:
rabbit MGCDSRKPLL LLPLLPLALV LQPW-SARGR ASAEEPSSIS PDKLLVITVA ... guineapig MAFGA----W LLLLPLLLLP PPPGACASDQ PRGSNP--VN PEKLLVITVA ... opossum ---------- ---------- ---------- ---------- SDKLLVITAA ... rat MAASVPEPRL LLLLLLLLPP LPPVTSASDR PRGANP--VN PDKLLVITVA ... elephant MASWGPGARL LLLLLLLLLP PPPATSASDR SRGSDR--VN PERLLVITVA ... human MTSSGPGPRF LLLLPLLL-- -PPAASASDR PRGRDP--VN PEKLLVITVA ... baboon MTSSRPGLRL LLLLLLL--- -PPAASASDR PRGRDP--VN PEKLLVMTVA ... dog MASSGPGLRL LLGLLLLL-P PPPATSASDR PRGGDP--VN PEKLLVITVA ... pig AMASGPGLR- LLLLPLLVLS PPPAASASDR PRGSDP--VN PDKLLVITVA ...
Phylogenetic tree in newick format:
((opossum:0.09636245,rabbit:0.85794020):0.05219782,(rat:0.07263127,elephant:0.03306863):0.01043531,(dog:0.01700528,(pig:0.02891345,(guineapig:0.14451043,(human:0.01169266,baboon:0.00827402):0.02619598):0.00816185):0.00631423):0.00800806);
- Step 3: The result of the previous step will be several trees, one for every group. Ideally, all trees would be identical, showing the real evolutionary history of the 9 species. But it is not easy to infer the real tree from sequence data, so trees from different groups might differ. Therefore, in the last step, we will build a consensus tree. This can be done by using an interactive tool called phylip.
- Output is a single consensus tree.
Files and submitting
Our goal for today is to build a pipeline that automates the whole task using make and execute it remotely using qsub. Most of the work is already done, only small modifications are necessary.
- Submit by copying requested files to /submit/hw03/username/
- Do not forget to submit protocol, outline of the protocol is in /tasks/hw03/protocol.txt
Start by copying /tasks/hw03 to your user directory
- cp -ipr /tasks/hw03 ~
It contains 3 subdirectories:
- large: larger sample of proteins for task A
- tiny: very small set of proteins for task B
- small: slightly larger set of proteins for task C
Task A
- In this task, you will run a long alignment job (>2 hours)
- Use directory large with files:
- ref.fa: selected human proteins
- other.fa: selected proteins from 8 other mammalian species
- Makefile: run blast on ref.fa vs other.fa (also formats database other.fa before that)
- run make -n to see what commands will be done (you should see makeblastdb and blastp + echo for timing), copy the output to the protocol
- run qsub with appropriate options to run make (at least -cwd and -b y)
- then run qstat > queue.txt
- Submit file queue.txt showing your job waiting or running
- When your job finishes, submit also the following two files:
- the last 100 lines from the output file ref.blast under the name ref-end.blast (use tool tail -n 100)
- standard output from the qsub job, which is stored in a file named e.g. make.oX where X is the number of your job. The output shows the time when your job started and finished (this information was written by commands echo in the Makefile)
Task B
- In this task, you will finish a Makefile for splitting blast results into ortholog groups and building phylogenetic trees for each group
- This Makefile works with much smaller files and so you can run it many times on vyuka, without qsub
- Work in directory tiny
- ref.fa: 2 human proteins
- other.fa: a selected subset of proteins from 8 other mammalian species
- Makefile: a longer makefile
- brm.pl: a Perl script for finding ortholog groups and sorting them to directories
The Makefile runs the analysis in four stages. Stages 1,2 and 4 are done, you have to finish stage 3
- If you run make without argument, it will attempt to run all 4 stages, but stage 3 will not run, because it is missing
- Stage 1: run as make ref.brm
- It runs blast as in task A, then splits proteins into ortholog groups and creates one directory for each group with file prot.fa containing protein sequences
- Stage 2: run as make alignments
- In each directory with a single gene, it will create an alignment prot.phy and link it under names lg.phy and wag.phy
- Stage 3: run as make trees (needs to be written by you)
- In each directory with a single gene, it should create lg.phy_phyml_tree and wag.phy_phyml_tree
- These corresponds to results of phyml commands run with two different evolutionary models WAG and LG, where LG is the default
- Run phyml by commands of the forms:
- phyml -i INPUT --datatype aa --bootstrap 0 --no_memory_check >LOG
- phyml -i INPUT --model WAG --datatype aa --bootstrap 0 --no_memory_check >LOG
- Change INPUT and LOG in the commands to appropriate filenames using make variables $@, $^, $* etc. Input should come from lg.phy or wag.phy in the directory of a gene and log should be the same as tree name with extension .log added (e.g. lg.phy_phyml_tree.log)
- Also add variables LG_TREES and WAG_TREES listing filenames of all desirable trees and uncomment phony target trees which uses these variables
- Stage 4: run as make consensus
- Output trees from stage 3 are concatenated for each model separately to files lg/intree, wag/intree and then phylip is run to produce consensus trees lg.tree and wag.tree
- This stage also needs variables LG_TREES and WAG_TREES to be defined by you.
- Run your Makefile
- Submit the whole directory tiny, including Makefile and all gene directories with tree files.
Task C
- Copy your Makefile from part B to directory small, which contains 9 human proteins and run make on this slightly larger set
- Again, run it on vyuka server without qsub, but it will take some time, particularly if the server is busy
- Look at the two trees from task C (wag.tree, lg.tree) using the figtree program on vyuka (you can also install it on your computer)
- In figtree, change the position of the root in the tree to make opossum the outgroup (species branching as the first away from the others).
- This is done in figtree by clicking on opossum and thus selecting it, then pressing Reroot button.
- Also switch on displaying branch labels. These labels show for each branch of the tree, how many of the input trees support this branch.
- Use the left panel with options.
- Export the trees in pdf format as wag.tree.pdf and lg.tree.pdf and include in your submission
- Compare the two trees and write your observations to the protocol
- Note that the two children of each internal node are equivalent, so their placement higher or lower in the figure does not matter.
- Do the two trees differ? What is the highest and lowest support for a branch in each tree?
- Also compare your trees with the accepted "correct tree" found here http://genome-euro.ucsc.edu/images/phylo/hg38_100way.png (note that this tree contains many more species, but all ours are included)
- Submit the entire small directory (including the two pdf files)
Further possibilities
Here are some possibilities for further experiments, in case you are interested (do not submit these):
- You could copy your extended Makefile to directory large and create trees for all ortholog groups in the big set
- This would take a long time, so submit it through qsub and only some time after the lecture is over to allow classmates to work on task A
- After ref.brm si done, programs for individual genes can be run in parallel, so you can try running make -j 2 and request 2 threads from qsub
- Phyml also supports other models, for example JTT (see manual), you could try to play with those.
- Command touch FILENAME will change modification time of the given file to current file
- What happens when you run touch on some of the intermediate files in the analysis in task B? Does Makefile always run properly?
L04
- Program for today: basics of Python and SQL
- Two version of homework: four easier tasks for beginners, or two more complicated ones for advanced Python/SQL programmers
- The next three lectures
- Computer science students will use Python and SQLite3 and several advanced Python libraries for complex data processing
- Bioinformatics students will use several bioinformatics command-line tools
Overview, documentation
Python: good sources for beginners:
SQL:
- Language for working with relational databases, more in a dedicated course
- We will cover basics of SQL and work with a simple DB system SQLite3
- SQLite3 documentation: [16]
- SQL tutorial: [17]
- SQLite3 in Python [18]
Program for today:
- We introduce a simple data set
- We look at several python scripts for processing this data set
- HW: You create another such script
- We introduce basics of working directly with SQLite3
- HW: You write your own queries
- We look at how to combine Python and SQLite
- HW: You write a program combining the two
Dataset for this week
- IMDb is an online database of movies and TV series with user ratings
- We have downloaded a preprocessed dataset of selected TV series ratings from GitHub
- From this dataset, we have selected only 10 series with the highest average number of voting users
- Data are 2 files in csv format: list of series, list of episodes
File series.cvs contains one row per series
- Columns: (0) series id, (1) series title, (2) TV channel:
3,Breaking Bad,AMC 2,Sherlock,BBC 1,Game of Thrones,HBO
File episodes.csv contains one row per episode:
- Columns: (0) series id, (1) episode title, (2) episode order within the whole series, (3) season number, (4) episode number within season, (5) user rating, (6) the number of votes
- Here is a sample of 4 episodes from Game of Thrones
- If the episode title contains a comma, the whole title is in quotation marks
1,"Dark Wings, Dark Words",22,3,2,8.6,12714 1,No One,58,6,8,8.3,20709 1,Battle of the Bastards,59,6,9,9.9,138353 1,The Winds of Winter,60,6,10,9.9,93680
Several python scripts
prog1.py
Print the second column (series title) from series.csv
#! /usr/bin/python3
# open a file for reading
with open('series.csv') as csvfile:
# iterate over lines of the input file
for line in csvfile:
# split a line into columns at commas
columns = line.split(",")
# print the second column
print(columns[1])
prog2.py
Print the list of series of each TV channel
- For illustration we also separately count the series for each channel, but the count could be obtained as the length of the list
- For simplicity we use library data structure defaultdict instead of plain python dictionary
#! /usr/bin/python3
from collections import defaultdict
# Create a dictionary in which default value
# for non-existent key is 0 (type int)
# For each channel we will count the series
channel_counts = defaultdict(int)
# Create a dictionary for keeping a list of series per channel
# default value empty list
channel_lists = defaultdict(list)
# open a file and iterate over lines
with open('series.csv') as csvfile:
for line in csvfile:
# strip whitespace (e.g. end of line) from end of line
line = line.rstrip()
# split line into columns, find channel and series names
columns = line.split(",")
channel = columns[2]
series = columns[1]
# increase counter for channel
channel_counts[channel] += 1
# add series to list for the channel
channel_lists[channel].append(series)
# print counts
print("Counts:")
for channel in channel_counts:
print("The number of series for channel \"%s\" is %d"
% (channel, channel_counts[channel]))
# print series lists
print("\nLists:")
for channel in channel_lists:
list = ", ".join(channel_lists[channel])
print("series for channel \"%s\": %s" % (channel,list))
prog3.py
Find the episode with the highest number of votes among all episodes
- We use a library for csv parsing to deal with quotation marks.
#! /usr/bin/python3
import csv
#keep maximum number of votes and its episode
max_votes = 0
max_votes_episode = None
# open a file
with open('episodes.csv') as csvfile:
# create a reader for parsing csv files
reader = csv.reader(csvfile, delimiter=',', quotechar='"')
# iterate over rows already split into columns
for row in reader:
votes = int(row[6])
if votes > max_votes:
max_votes = votes
max_votes_episode = row[1]
# print result
print("Maximum votes %d in episode \"%s\"" % (max_votes, max_votes_episode))
prog4.py
Example of function definition, reading the whole file into a 2d array
#! /usr/bin/python3
import csv
def read_csv_to_list(filename):
# create empty list
rows = []
# open a file
with open(filename) as csvfile:
# create a reader for parsing csv files
reader = csv.reader(csvfile, delimiter=',', quotechar='"')
# iterate over rows already split into columns
for row in reader:
rows.append(row)
return rows
series = read_csv_to_list('series.csv')
episodes = read_csv_to_list('episodes.csv')
print("the number of episodes is %d" % len(episodes))
# further processing of series and episodes...
Now do #HW04, task A
SQL and SQLite
Creating a database
SQLite3 database is a file with your data stored in some special format. To load our csv file to a SQLite database, run command:
sqlite3 series.db < create_db.sql
Contents of create_db.pl:
CREATE TABLE series ( id INT, title TEXT, channel TEXT ); .mode csv .import series.csv series CREATE TABLE episodes ( seriesId INT, title TEXT, orderInSeries INT, season INT, orderInSeason INT, rating REAL, votes INT ); .mode csv .import episodes.csv episodes
SQL queries
- Run sqlite3 series.db
- Then type on SQLite3 command line the following queries
/* switch on human-friendly formatting */ .mode column .headers on /* print title of each series (as prog1.py) */ SELECT title FROM series; /* sort titles alphabetically */ SELECT title FROM series ORDER BY title; /* find the highest vote number among episodes */ SELECT MAX(votes) FROM episodes; /* find the episode with the highest number of votes, as prog3.py */ SELECT title, votes FROM episodes ORDER BY votes DESC LIMIT 1; /* print all episodes with at least 50k votes, order by votes */ SELECT title, votes FROM episodes WHERE votes>50000 ORDER BY votes desc; /* join series and episodes tables, print 10 episodes * with the highest number of votes */ SELECT s.title, e.title, votes FROM episodes AS e, series AS s WHERE e.seriesId=s.id ORDER BY votes desc limit 10; /* compute the number of series per channel, as prog2.py */ SELECT channel, COUNT() as series_count FROM series GROUP BY channel; /* print the number of episodes and average rating per season and series */ SELECT seriesId, season, COUNT() AS episode_count, AVG(rating) AS rating FROM episodes GROUP BY seriesId, season;
Now do #HW04, tasks B1, B2
Accessing a database from Python
read_db.py
- Script illustrates running a SELECT query and getting results
#! /usr/bin/python3
import sqlite3
# connect to a database
connection = sqlite3.connect('series.db')
# create a "cursor" for working with the database
cursor = connection.cursor()
# run a select query
# supply parameters of the query using placeholders ?
threshold = 40000
cursor.execute("""SELECT title, votes FROM episodes
WHERE votes>? ORDER BY votes desc""", (threshold,))
# retrieve results of the query
for row in cursor:
print("Episode \"%s\" votes %s" % (row[0],row[1]))
# close db connection
connection.close()
write_db.py
Script illustrates creating a new database containing a multiplication table
#! /usr/bin/python3
import sqlite3
# connect to a database
connection = sqlite3.connect('multiplication.db')
# create a "cursor" for working with the database
cursor = connection.cursor()
cursor.execute("""
CREATE TABLE mult_table (
a INT, b INT, mult INT)
""")
for a in range(1,11):
for b in range(1,11):
cursor.execute("INSERT INTO mult_table (a,b,mult) VALUES (?,?,?)",
(a,b,a*b))
# important: save the changes
connection.commit()
# close db connection
connection.close()
We can check the result by running command
sqlite3 multiplication.db "SELECT * FROM mult_table;"
Now do #HW04, task C
HW04
Introduction
Choose one of the options:
- Tasks A, B1, B2, C (recommended for beginners)
- Tasks C, D (recommended for experienced Python/SQL programmers)
Preparation
Copy files:
mkdir hw04 cd hw04 cp -iv /tasks/hw04/* .
The directory contains the following files:
- *.py: python scripts from the lecture, included only for convenience
- series.csv, episodes.csv: data file used in the homework (and the lecture)
- create_db.sql: sql commands to create the database needed in tasks B1, B2, C, D
- protocol.txt: fill in and submit the protocol. Only "Vyhodnotenie" and "Pouzite zdroje" are needed this time
Task A
- Write a script which reads both csv files and outputs for each TV channel the total number of episodes in their series combined
- Submit file taskA.py with your script
- Run your script as follows and submit the file taskA.txt:
./taskA.py > taskA.txt
- One of the lines of your output should be:
The number of episodes for channel "HBO" is 76
Hints:
- A good place to start is prog4.py with reading both csv files and prog2.py with a dictionary of counters
- It might be useful to build a dictionary linking the series id to the channel name for that series
Task B1
- To prepare the database for tasks B1, B2 and C, run the command:
sqlite3 series.db < create_db.sql
To verify that your database was created correctly, you can run the following commands:
sqlite3 series.db ".tables" # output should be episodes series sqlite3 series.db "select count() from episodes; select count() from series;" # output should be 348 and 10
- The last query in the lecture counts the number of episodes and average rating per each season of each series:
SELECT seriesId, season, COUNT() AS episode_count, AVG(rating) AS rating FROM episodes GROUP BY seriesId, season;
- Use join with the series table to replace the numeric series id with the series title and add the channel name
- Write your SQL query to file taskB1.sql and submit this file
- The first two lines of the sql file should be
.mode column .headers on
- Run your query as follows:
sqlite3 series.db < taskB1.sql > taskB1.txt
- Submit also the resulting file taskB1.txt
- For example, both seasons of True Detective by HBO have 8 episodes and average ratings 9.3 and 8.25
True Detective HBO 1 8 9.3 True Detective HBO 2 8 8.25
Task B2
- For each channel compute the total count and average rating of all their episodes.
- Write your SQL query to file taskB2.sql and submit this file
- The first two lines of the sql file should be
.mode column .headers on
- Run your query as follows:
sqlite3 series.db < taskB2.sql > taskB2.txt
- Submit also the resulting file taskB2.txt
- For example, all 76 episodes for the two HBO series have average rating as follows:
HBO 76 8.98947368421053
Task C
- If you have not done so already, create an SQLite database, as explained at the beginning of task B1.
- Write a python script that runs the last query from the lecture (shown below) and stores its results in a separate table called seasons in the series.db database
/* print the number of episodes and average rating per season and series */ SELECT seriesId, season, COUNT() AS episode_count, AVG(rating) AS rating FROM episodes GROUP BY seriesId, season;
- SQL can store results from a query directly in a table, but in this task you should instead read each row of the SELECT query in python and to store it by running INSERT command from python
- Also do not forget to create the new table in the database with appropriate column names and types. You can execute CREATE TABLE command from python
- The cursor from the SELECT query is needed while you iterate over the results. Therefore create two cursors - one for reading the database and one for writing.
- If you change your database during debugging, you can start over by running the command for creating the database above
- Store and submit the script in taskC.py. Also submit the modified database series.db
- To check that your table was created, you can run command
- sqlite3 series.db "SELECT * FROM seasons;"
- This will print many lines, including this one: "5|1|8|9.3" which is for season 1 of series 5 (True Detective)
Task D
- For each pair of consecutive seasons within each series, compute how much has the average rating increased or decreased
- For example in the Sherlock series, season 1 had rating 8.825 and season 2 rating 9.26666666666667, and thus the difference in ratings is -0.44166666666667
- Print a table containing series name, season number x, average rating in season x and average rating in season x+1
- The table should be ordered by the difference between the last two columns, i.e. from seasons with the highest increase to seasons to the highest drop.
- One option is to run a query in SQL in which you join table seasons from task C with itself and select rows that belong to the same series and successive seasons
- You can also read the rows of the seasons table in Python, combine information from rows for successive seasons of the same series and create the final report by sorting
- Submit your code as taskD.py or taskD.sql
- Submit the resulting table as taskD.txt
The output should start like this (the formatting may differ):
series season x rating for x rating for x+1 ---------- ---------- ------------ ---------------- Sherlock 1 8.825 9.26666666666667 Breaking B 4 9.0 9.375
When using sql without python, include the following two lines in taskD.sql
.mode column .headers on
and run your query as sqlite3 series.db < taskD.sql > taskD.txt
L05inf
In this lecture we dive into SQLite3 and Python.
SQLite3
SQLite3 is a simple "database" stored in one file. Think of SQLite not as a replacement for Oracle but as a replacement for fopen(). Documentation: https://www.sqlite.org/docs.html
You can access sqlite database either from command line:
usamec@Darth-Labacus-2:~$ sqlite3 db.sqlite3 SQLite version 3.8.2 2013-12-06 14:53:30 Enter ".help" for instructions Enter SQL statements terminated with a ";" sqlite> CREATE TABLE test(id integer primary key, name text); sqlite> .schema test CREATE TABLE test(id integer primary key, name text); sqlite> .exit
Or from python interface: https://docs.python.org/2/library/sqlite3.html.
Python
Python is a perfect language for almost anything. Here is a cheatsheet: http://www.cogsci.rpi.edu/~destem/igd/python_cheat_sheet.pdf
Scraping webpages
The simplest tool for scraping webpages is urllib2: https://docs.python.org/2/library/urllib2.html Example usage:
import urllib2
f = urllib2.urlopen('http://www.python.org/')
print f.read()
Or use requests package:
import requests
r = requests.get("http://en.wikipedia.org")
print(r.text[:10])
Parsing webpages
We use beautifulsoup4 for parsing html (http://www.crummy.com/software/BeautifulSoup/bs4/doc/). I recommend following examples at the beginning of the documentation and example about CSS selectors: http://www.crummy.com/software/BeautifulSoup/bs4/doc/#css-selectors
Parsing dates
You have two options. Either use datetime.strptime or use dateutil package.
Other usefull tips
- Don't forget to commit to your sqlite3 database (db.commit()).
- CREATE TABLE IF NOT EXISTS can be usefull at the start of your script.
- Inspect element (right click on element) in Chrome can be very helpful.
- Use screen command for long running scripts.
- All packages are installed on vyuka server. If you are planning using your own laptop, you need to install them using pip (preferably using virtualenv).
HW05inf
- Submit by copying requested files to /submit/hw05inf/username/
General goal: Scrape comments from several (hundreds) sme.sk users from last month and store them in SQLite3 database.
Task A
Create SQLite3 "database" with appropriate schema for storing comments from SME.sk discussions. You will probably need tables for users and comments. You don't need to store which comments replies to which one.
Submit two files:
- db.sqlite3 - the database
- schema.txt - brief description of your schema and rationale behind it
Task B
Build a crawler, which crawls comments in sme.sk discussions. You have two options:
- For fewer points: Script which gets url of the user (http://ekonomika.sme.sk/diskusie/user_profile.php?id_user=157432) and crawls his comments from last month.
- For more points: Scripts which gets one starting url (either user profile or some discussion, your choice) and automatically discovers users and crawls their comments.
This crawler should store comments in SQLite3 database built in previous task. Submit following:
- db.sqlite3 - the database
- every python script used for crawling
- README (how to start your crawler)
L05bin
The goal of the next three lectures is to get experience with several common bioinformatics tools
- You will learn more about the algorithms and models behind these tools in Methods in bioinformatics course
Overview of DNA sequencing and assembly
- DNA sequencing is a technology of reading the order of nucleotides along a DNA strand
- The result is represented as a string of A,C,G,T
- Only fragments of DNA of limited length can be read, these are called sequencing reads
- Different technologies produce reads of different characteristics
- Examples:
- Illumina sequencers produce short reads (typical length 100-200bp), but in great quantities and very low error rate (<0.1%)
- The reads usually come in pairs sequenced from both ends of a DNA fragment of an approximately known length
- Oxford nanopore sequencers produce longer reads (thousands bp or more), but the error rates are higher (10-15%)
- Illumina sequencers produce short reads (typical length 100-200bp), but in great quantities and very low error rate (<0.1%)
- The goal of genome sequencing is to read all chromosomes of an organism
- Sequencing machines produce many reads coming from different parts of the genome
- Using software tools called sequence assemblers, these reads are glued together based on overlaps
- Ideally we would get the true chromosomes, but often we get only shorter fragments called contigs
- The results of assembly can contain errors
- We prefer longer contigs with fewer errors
Sequence alignments and dotplots
- Sequence alignment is the task of finding similarities between DNA (or protein) sequences
- Here is an example - short similarity between region 344447..344517 of one sequence and 3261..3327 of another
Query: 344447 tctccgacggtgatggcgttgtgcgtcctctatttcttttatttctttttgttttatttc 344506
|||||||| |||||||||||||||||| ||||||| |||||||||||| || ||||||
Sbjct: 3261 tctccgacagtgatggcgttgtgcgtc-tctatttattttatttctttgtg---tatttc 3316
Query: 344507 tctgactaccg 344517
|||||||||||
Sbjct: 3317 tctgactaccg 3327
- Alignments can be stored in many formats and visualised as dotplots
- In a dotplot, x-axis correspond to positions in one sequence and y-axis to another sequence
- Diagonal lines show alignments between the sequences (direction of the diagonal shows which DNA strand was aligned)
File formats
Fasta
- For storing DNA, RNA and protein sequences
- We were already working with fasta on #HW01
- Each sequence consists of several lines of the file. The first line starts with ">" followed by identifier of the sequence and optionally some further description separated by whitespace
- The sequence itself is on the second line, long sequences are split into multiple lines
>SRR022868.1845_1 AAATTTAGGAAAAGATGATTTAGCAACATTTAGCCTTAATGAAAGACCAGATTCTGTTGCCATGTTTGAATGCCTTAAACCAGTAGCAGAATCAGTATAAA >SRR022868.1846_1 TAGCGTTGTAAAATAAATTTCTAGAATGGAAGTGATGATATTGAAATACACTCAGATCCTGAATGAAAGATTTATTAAAGTTAAGACGAGAGTCTCATTAT
Fastq
- Special format for storing sequencing reads, containing DNA sequences but also quality information about each nucleotide * More in Lecture 01
Sam/bam
- Format for storing alignments of sequencing reads (or other sequences) to a genome [19]
- For each read, the file contains the read itself, its quality, but also the chromosome/contig name and position where this read is likely coming from, and an additional information e.g. about mapping quality (confidence in the correct location)
- Sam files are text-based, thus easier to check manually; bam files are binary and compressed, thus smaller and faster to read
- We can easily convert between them using samtools
Paf format
- Another format for storing alignments [20]
Gzip
- A general-purpose format for file compression [21]
- Often used in bioinformatics on large fastq or fasta files
- Running command gzip filename.ext will create compressed file filename.ext.gz (original file will be deleted).
- The reverse process by gunzip filename.ext.gz
- Print the content of a gzipped file zcat filename.ext.gz (this will keep the gzipped file as is)
- Page through the content of a gzipped file zless filename.ext.gz
HW05bin
Submit the protocol and the required files to /submit/hw05bin
Task A: examine input files
- copy files from /tasks/hw05bin/
mkdir hw05 cd hw05 cp -iv /tasks/hw05bin/* .
- ref.fasta is a piece of genome from E.coli
- miseq_R1.fastq.gz and miseq_R2.fastq.gz are sequencing reads from Illumina MiSeq sequencer. First reads in pairs are in R1 file, second reads in R2 file. These reads come from the region in ref.fasta
- nanopore.fasta are nanopore sequencing reads in a fasta format (without qualities). These reads are also from the region in ref.fasta
Try to find the answers to the following questions using command-line tools. In your protocol, note down the commands as well as the answers:
- (a) How many reads are in the miseq files? Is the number of reads the same in both files?
- Try command zcat miseq_R1.fastq.gz | wc -l
- Can you figure out the answer from the result of this command?
- (b) How long are individual reads in the miseq files?
- Look at the file using zless - do all reads appear to be of an equal length?
- Extend the following command with tail and wc -c to get the length of the first read: zcat miseq_R1.fastq.gz | head -n 2
- Repeat for both miseq files
- (c) How many reads are in the nanopore file (beware - different format)
- (d) What is the average length of the reads in the nanopore file?
- Try command: samtools faidx nanopore.fasta
- This creates nanopore.fasta.fai file, where the second column contains sequence length of each read
- Compute the average of this column by a one-liner: perl -lane '$s+=$F[1]; $n++; END { print $s/$n }' nanopore.fasta.fai
- (e) How long is the sequence in the ref.fasta file?
Task B: assemble the sequence from the reads
- We will pretend that the correct answer (ref.fasta) is not known and we will try to assemble it from the reads
- We will assemble Illumina reads by program SPAdes and nanopore reads by miniasm
- Assembly takes several minutes, we will run it in the background using screen command
SPAdes
- Run screen -S spades
- Press Enter to get command-line, then run the follwing command:
- spades.py -t 1 -m 1 --pe1-1 miseq_R1.fastq.gz --pe1-2 miseq_R2.fastq.gz -o spades > spades.log
- Press Ctrl-a followed by d
- This will take you out of screen command
- Run top command to check that your command is running
Minimap
- Create file minimap.sh containing the following commands:
# Find alignments between pairs of reads minimap2 -x ava-ont -t 1 nanopore.fasta nanopore.fasta | gzip -1 > nanopore.paf.gz # Use overlaps to compute assembled genome miniasm -f nanopore.fasta nanopore.paf.gz > miniasm.gfa 2> miniasm.log # Convert genome to fasta format perl -lane 'print ">$F[1]\n$F[2]" if $F[0] eq "S"' miniasm.gfa > miniasm.fasta # Align reads to the assembled genome minimap2 -x map-ont --secondary=no -t 1 miniasm.fasta nanopore.fasta | gzip -1 > miniasm.paf.gz # Polish the genome by finding consensus of aligned reads at each position racon -t 1 -u nanopore.fasta miniasm.paf.gz miniasm.fasta > miniasm2.fasta
- Run screen -S minimap
- In screen, run source ./minimap.sh
- Press Ctrl-a d to exit screen
To check if your commands have finished:
- Re-enter the screen environment using screen -r spades or screen -r miniasm
- If the command finished, terminate screen by pressing Ctrl-d or typing exit
Examine the outputs, write commands and answers to your protocol:
- Copy output of spades under a new filename: cp -ip spades/contigs.fasta spades.fasta
- Output of miniasm should be in miniasm2.fasta
- (a) How many contigs are in each of these two files?
- (b) What can you find out from the names of contigs in spades.fasta? What is the length of the shortest and longest contigs? Cov in the names is abbreviation of read coverage - the average number of reads covering a position on the contig. Do the reads have similar coverage, or are there big differences?
- Use command grep '>' spades.fasta
- (c) What are the lengths of contigs in miniasm2.fa file? (you can use LN:i: in the name of contigs)
Submit files miniasm2.fasta and spades.fasta
Task C: compare assemblies using Quast command
- We have found basic characteristics of the two assemblies in task B
- Now we will use program Quast to compare both assemblies to the correct answer in ref.fa
quast.py -R ref.fasta miniasm2.fasta spades.fasta -o stats
- Submit file stats/report.txt
Look at the results in stats/report.txt and answer the following questions in your protocol:
- (a) How many contigs quast reported in the two assemblies? Does it agree with your counts in part B?
- (b) What is the number of mismatches per 100kb in the two assemblies? Which one is better? Why do you think it is so? (look at the properties of used sequencing technologies in the lecture)
- (c) What portion of the reference sequence is covered by the two assemblies (genome fraction)? Which assembly is better in this aspect?
- (d) What is the length of the longest alignment between contigs and the reference in the two assemblies? Which assembly is better in this aspect?
Task D: create dotplots of assemblies
- We will now visualize alignments between each assembly and the reference genome using dotplots
- As in other tasks, write commands and answers to your protocol
- (a) Create dotplot comparing miniasm assembly to the reference sequence
# alignments minimap2 -x asm10 -t 1 ref.fasta miniasm2.fasta > ref-miniasm2.paf # creating dotplot /usr/local/share/miniasm/miniasm/minidot -f 12 ref-miniasm2.paf | ps2pdf -dEPSCrop - ref-miniasm2.pdf # displaying dotplot - if this does not work, copy the pdf file to your commputer and view there evince ref-miniasm2.pdf &
- x-axis is reference, y-axis assembly
- Which part of the reference is missing in the assembly?
- Do you see any other big differences between the assembly and the reference?
- (b) Use analogous commands to create dotplot for spades assembly, call it ref-spades.pdf
- What are vertical gray lines in the dotplot?
- Is any contig aligning to multiple places in the reference? To how many places?
- (c) Use analogous commands to create dotplot of reference to itself, call it ref-ref.pdf
- However, in the minimap2 command add option -p 0 to include also weaker self-alignments
- Do you see any self-alignments, showing repeated sequences in the reference? Does this agree with dotplot in part (b)?
- Submit all three pdf files ref-miniasm2.pdf, ref-spades.pdf, ref-ref.pdf
Task E: Align reads and assemblies to reference, visualize in igv
- Finally, we will align all source reads as well as assemblies to the reference genome, then visualize alignment in igv tool
- Write commands and answers to your protocol
- Submit all four bam files ref-miseq.bam, ref-nanopore.bam, ref-spades.bam, ref-miniasm2.bam
- (a) Align illumina reads (miseq files) to reference sequence
# align illumina reads to reference # minimap produces sam file, samtools view converts to bam, samtools sort orders by coordinate minimap2 -a -x sr --secondary=no -t 1 ref.fasta miseq_R1.fastq.gz miseq_R2.fastq.gz | samtools view -S -b - | samtools sort - ref-miseq # index bam file for faster access samtools index ref-miseq.bam
- (b) Similarly align nanopore reads. but instead of -x sr use -x map-ont, call the result ref-nanopore.bam, ref-nanopore.bam.bai
- (c) Similarly align spades.fasta, but instead of -x sr use -x asm10, call the result ref-spades.bam
- (d) Similarly align miniasm2.fasta, but instead of -x sr use -x asm10, call the result ref-miniasm2.bam
- (e) Run igv viewer. Beware: It needs a lot of memory, do not keep open unnecessarily
- igv -g ref.fasta &
- Using Menu->File->Load from File open all bam four files
- Look at region ecoli-frag:224,000-244,000
- How many spades contigs do you see aligning in this region?
- Look at region ecoli-frag:227,300-227,600
- Try to comment what you see, how frequent are errors in individual assemblies and read sets?
- If you are unable to run igv from home, you can install it on your computer [22] and download ref.fasta and all bam and .bam.bai files
L06inf
In this lecture we will use Flask and simple text processing utilities from ScikitLearn.
Flask
Flask is simple web server for python (http://flask.pocoo.org/docs/1.0/quickstart/) You can find sample flask application at /tasks/hw06/simple_flask.
You can run it using these commands:
cd <your directory> export FLASK_APP=main.py export FLASK_ENV=development (this is optional, but recommended for debugging) flask run --host=0.0.0.0 --port=4247 (on vyuka server this starts python2.7, if you want python3 use flask3 run, but that is only on vyuka, on your own computer use virtualenv)
Before running change the port number. You can then access your app at vyuka.compbio.fmph.uniba.sk:4247 (change port number).
There may be problem with access to strange port numbers due to firewalling rules. There are at least two ways to circumvent this:
- Use X forwarding and run web browser directly from vyuka
local_machine> ssh vyuka.compbio.fmph.uniba.sk -XC vyuka> chromium-browser
- Create SOCKS proxy to vyuka.compbio.fmph.uniba.sk and set SOCKS proxy at that port on your local machine. Then all web traffic goes through vyuka.compbio.fmph.uniba.sk via ssh tunnel. To create SOCKS proxy server on local machine port 8000 to vyuka.compbio.fmph.uniba.sk:
local_machine> ssh vyuka.compbio.fmph.uniba.sk -D 8000
(keep ssh session open while working)
Flask uses jinja2 (http://jinja.pocoo.org/docs/dev/templates/) templating language for showing html (you can use strings in python but it is painful).
Processing text
Main tool for processing text is CountVectorizer class from ScikitLearn (http://scikit--learn.org/stable/modules/generated/sklearn.feature_extraction.text.CountVectorizer.html). It transforms text into bag of words (for each word we get counts). Example:
from sklearn.feature_extraction.text import CountVectorizer vec = CountVectorizer(strip_accents='unicode') texts = [ "Ema ma mamu.", "Zirafa sa vo vani kupe a hneva sa." ] t = vec.fit_transform(texts).todense() print(t) print(vec.vocabulary_)
Useful things
We are working with numpy arrays here (that's array t in example above) Numpy arrays has also lots of nice tricks. First lets create two matrices:
>>> import numpy as np
>>> a = np.array([[1,2,3],[4,5,6]])
>>> b = np.array([[7,8],[9,10],[11,12]])
>>> a
array([[1, 2, 3],
[4, 5, 6]])
>>> b
array([[ 7, 8],
[ 9, 10],
[11, 12]])
We can sum this matrices or multiply them by some number:
>>> 3 * a
array([[ 3, 6, 9],
[12, 15, 18]])
>>> a + 3 * a
array([[ 4, 8, 12],
[16, 20, 24]])
We can calculate sum of elements in each matrix, or sum by some axis:
>>> np.sum(a) 21 >>> np.sum(a, axis=1) array([ 6, 15]) >>> np.sum(a, axis=0) array([5, 7, 9])
There is a lot other useful functions check https://docs.scipy.org/doc/numpy-dev/user/quickstart.html.
This can help you get top words for each user: http://docs.scipy.org/doc/numpy/reference/generated/numpy.argsort.html#numpy.argsort
HW06inf
- Submit by copying requested files to /submit/hw06inf/username/
General goal: Build a simple website, which lists all crawled users and for each users has a page with simple statistics for given user.
This lesson requires crawled data from previous lesson, if you don't have one, you can find it at (and thank Baska): /tasks/hw06inf/db.sqlite3
Submit source code (web server and preprocessing scripts) and database files.
Task A
Create a simple flask web application which:
- Has a homepage where is a list of all users (with links to their pages).
- Has a page for each user, which has simple information about user: His nickname, number of posts and hist last 10 posts.
Task B
For each user preprocess and store list of his top 10 words and list of top 10 words typical for him (which he uses much more often than other users, come up with some simple heuristics). Show this information on his page.
Task C
Preprocess and store list of top three similar users for each user (try to come up with some simple definition of similarity based on text in posts). Again show this information on user page.
Bonus: Try to use some simple topic modeling (e.g. PCA as in TruncatedSVD from scikit-learn) and use it for finding similar users.
L06bin
Eukaryotic gene structure
- Recall the Central dogma of molecular biology: the flow of genetic information from DNA to RNA to protein (gene expression)
- In eukaryotes, mRNA often undergoes splicing, where introns are removed and exons are joined together
- The very start and end of mRNA remain untranslated (UTR = untranslated region)
- The coding part of the gene starts with a start codon, contains a sequence of additional codons and ends with a stop codon. Codons can be interrupted by introns.
Computational gene finding
- Input: DNA sequence (an assembled genome or a part of it)
- Output: positions of protein coding genes and their exons
- If we know the exact position of coding regions of a gene, we can use genetic code to predict the protein sequence encoded by it
- Gene finders use statistical features observed from known genes, such as typical sequence motifs near the start codons, stop codons and splice sites, typical codon frequences, typical exon and intron lengths etc.
- These statistical parameters need to be adjusted for each genome.
- We will use a gene finder called Augustus
Gene expression
- Not all genes undergo transcription and translation all the time and at the same level
- The processes of transcription and translation are regulated according to cell needs
- The term "gene expression" has two meanings
- the process of transcription and translation (synthesis of a gene product)
- the amount of mRNA or protein produced from a single gene (genes with high or low expression)
- RNA-seq technology can sequence mRNA extracted from a sample of cells
- We can aligned sequenced reads back to the genome
- The number of reads coming from a gene depends on its expression level (and on its length)
HW06bin
Input files, submitting
Copy files from /tasks/hw06bin/
mkdir hw06 cd hw06 cp -iv /tasks/hw06bin/* .
Files:
- ref.fasta is a 38kb piece of genome of the fungus Aspergillus nidulans
- rnaseq.fastq are RNA-seq reads from Illumina sequencer extracted from the Short read archive
- annot.gff is the reference gene annotation from the database (we will consider his as correct gene positions)
Submit the protocol and the required files to /submit/hw06bin
Task A: Gene finding
Run the Augustus gene finder with two versions of parameters:
- one trained specifically for A. nidulas genes
- one trained for the human genome, where genes have different statistical properties (for example, they are longer and have more introns)
augustus --species=anidulans ref.fasta > augustus-anidulans.gtf augustus --species=human ref.fasta > augustus-human.gtf
- The results of gene finding are in the GTF format. Rows starting with # are comments, each of the remaining rows describes some interval of the sequence. If the second column is CDS, it is a coding part of an exon.
- The reference annotation annot.gff is in a similar format called GFF3.
Examine the files and try to find the answers to the following questions using command-line tools
- (a) How many exons are in each of the two gtf files? (Beware: simply using grep with pattern CDS may yield lines containing this string in a different column. You can use e.g. techniques from #L02 and #HW02).
- (b) How many genes are in each of the two gtf files? (The files contain rows with word gene in the second column, one for each gene)
- (c) How many exons and genes are in the annot.gff file?
Write the anwsers and commands to the protocol. Submit files augustus-anidulans.gtf and augustus-human.gtf.
Task B: Aligning RNA-seq reads
- Align RNA-seq reads to the genome
- We will use a specialized tool tophat, which can recognize introns
- The we will sort and index bam the file, similarly as in #HW05bin
bowtie2-build ref.fasta ref.fasta tophat2 -i 10 -I 10000 --max-multihits 1 --output-dir rnaseq ref.fasta rnaseq.fastq samtools sort rnaseq/accepted_hits.bam rnaseq samtools index rnaseq.bam
In addition to the bam file, TopHat produced several other files in the rnaseq folder. Examine them to find out answers to the following questions (you can do it manually by looking at the the files, e.g. by less command):
- (a) How many reads were in the the fastq file? How many of them were successfully mapped?
- (b) How many introns ("junctions") were predicted? How many of them are supported by more than one read? (The 5th column of the corresponding file is the number of reads supporting a junction.)
Write anwsers to the protocol. Submit file rnaseq.bam.
Task C: Visualizing in igv
As before, run igv as follows:
igv -g ref.fasta &
- Open additional files using menu File -> Load from File
- annot.gff, augustus-anidulans.gtf, augustus-human.gtf, rnaseq.bam
- Exons are shown as thicker boxes, introns are thinner.
- For each of the following questions, select part of the sequence illustrating the answer and export figure using File->Save image
- You can check these images using command eog
Questions:
- (a) Create image illustrating differences between Augustus with human parameters and the reference annotation, save as a.png. Briefly describe the differences in words.
- (b) Find some differences between Augustus with A.nidulans parameters and the reference annotation. Store an illustrative figure as b.png. Which parameters have yielded a more accurate prediction?
- (c) Zoom in to one of the genes with high expression level and try to find spliced read alignments supporting the annotated intron boundaries. Store the image as c.png.
Submit files a.png, b.png, c.png. Write answers to your protocol.
L07inf
In this lesson we make simple javascript visualizations.
Your goal is to take examples from here https://developers.google.com/chart/interactive/docs/ and tweak them for your purposes.
Tips:
- You can output your data into javascript data structures in Flask template. It is a bad practice, but sufficient for this lesson. (Better way is to load JSON through API).
- Remember that you have to bypass the firewall.
HW07inf
- Submit by copying requested files to /submit/hw07inf/username/
General goal: Extend user pages from previous project with simple visualizations.
Task A
Show a calendar, which shows during which days was user active (like this https://developers.google.com/chart/interactive/docs/gallery/calendar#overview).
Task B
Show a histogram of comments length (like this https://developers.google.com/chart/interactive/docs/gallery/histogram#example).
Task C
Try showing a word tree for a user (https://developers.google.com/chart/interactive/docs/gallery/wordtree#overview). Try to normalize the text (lowercase, remove accents). CountVectorizer has method build_analyzer, which returns a function, which does this for you.
L07bin
Polymorphisms
- Individuals within species differ slightly in their genomes
- Polymorphisms are genome variants which are relatively frequent in a population (e.g. at least 1%)
- SNP: single-nucleotide polymorphism (a polymorphism which is a single substitution)
- Recall that most human cells are diploid, with one set of chromosomes inherited from the mother and the other from the father
- At a particular location, a single human can thus have two different alleles (heterozygosity) or two copies of the same allele (homozygosity)
Finding polymorphisms / genome variants
- We compare sequencing reads coming from an individual to a reference genome of the species
- First we align them, as in #HW05bin
- Then we look for positions where a substantial fraction of reads does not agree with the reference (SNP-calling)
Programs and file formats
- For mapping, we will use bwa mem (you can also try minimap2, as in #HW05bin)
- For SNP calling, we will use freebayes
- For reads and read alignments, we will use fastq and bam files, as in previous lectures
- For storing found variants, we will use VCF files
- For storing genome intervals, we will use BED files
Human variants
- For many human SNPs we already know something about their influence on phenotype and their prevalence in different parts of the world
- There are various databases, e.g. dbSNP, OMIM, or user-editable SNPedia
UCSC genome browser
- On-line tool similar to IGV
- http://genome-euro.ucsc.edu/
- Nice interface for browsing genomes, lot of data for some genomes (particularly human), but not all sequenced genomes represented
Basics
- on the front page, choose Genomes in the top blue menu bar
- select a genome and its version, optionally enter position or keyword, press submit
- on the browser screen top image shows chromosome map, selected region in red
- below a view of selected region and various track with information about this region
- for example some of the top tracks display genes (boxes are exons, lines are introns)
- tracks can be switched on and off and configured in the bottom part of the page
- different display levels, full contains all information but takes a lot of vertical space
- navigation at the top (move, zoom, etc.)
- various actions in the menu
- clicking at the browser figure allows you to get more information about a gene or other displayed item
- this week, we will need tracks GENCODE and dbSNP - check e.g. gene ACTN3 and within it SNP rs1815739 in exon15
Blat
- UCSC genome browser uses a fast but less sensitive BLAT (good for the same or very closely related species)
- Choose Tools->Blat in the top blue menu bar, enter DNA sequence below, search in the human genome
- What is the identity level for the top found match? What is its span in the genome? (Notice that other matches are much shorter)
- Using Details link in the left column you can see the alignment itself, Browser link takes you to the browser at the matching region
AACCATGGGTATATACGACTCACTATAGGGGGATATCAGCTGGGATGGCAAATAATGATTTTATTTTGAC TGATAGTGACCTGTTCGTTGCAACAAATTGATAAGCAATGCTTTCTTATAATGCCAACTTTGTACAAGAA AGTTGGGCAGGTGTGTTTTTTGTCCTTCAGGTAGCCGAAGAGCATCTCCAGGCCCCCCTCCACCAGCTCC GGCAGAGGCTTGGATAAAGGGTTGTGGGAAATGTGGAGCCCTTTGTCCATGGGATTCCAGGCGATCCTCA CCAGTCTACACAGCAGGTGGAGTTCGCTCGGGAGGGTCTGGATGTCATTGTTGTTGAGGTTCAGCAGCTC CAGGCTGGTGACCAGGCAAAGCGACCTCGGGAAGGAGTGGATGTTGTTGCCCTCTGCGATGAAGATCTGC AGGCTGGCCAGGTGCTGGATGCTCTCAGCGATGTTTTCCAGGCGATTCGAGCCCACGTGCAAGAAAATCA GTTCCTTCAGGGAGAACACACACATGGGGATGTGCGCGAAGAAGTTGTTGCTGAGGTTTAGCTTCCTCAG TCTAGAGAGGTCGGCGAAGCATGCAGGGAGCTGGGACAGGCAGTTGTGCGACAAGCTCAGGACCTCCAGC TTTCGGCACAAGCTCAGCTCGGCCGGCACCTCTGTCAGGCAGTTCATGTTGACAAACAGGACCTTGAGGC ACTGTAGGAGGCTCACTTCTCTGGGCAGGCTCTTCAGGCGGTTCCCGCACAAGTTCAGGACCACGATCCG GGTCAGTTTCCCCACCTCGGGGAGGGAGAACCCCGGAGCTGGTTGTGAGACAAATTGAGTTTCTGGACCC CCGAAAAGCCCCCACAAAAAGCCG
HW07bin
Input files, submitting
Copy files from /tasks/hw07bin/
mkdir hw07 cd hw07 cp -iv /tasks/hw07bin/* .
Files:
- humanChr7Region.fasta is a 7kb piece of the human chromosome 7
- motherChr7Region.fastq is a sample of reads from an anonymous donor known as NA12878, these reads come from region in humanChr7Region.fasta
- fatherChr12.vcf and motherChr12.vcf are single-nucleotide variants in chr12 obtained by sequencing two individuals NA12877, NA12878 (these come from a larger family)
Submit the protocol and the required files to /submit/hw07bin
Task A: read mapping and SNP calling
Align reads to reference:
bwa index humanChr7Region.fasta bwa mem humanChr7Region.fasta motherChr7Region.fastq | samtools view -S -b - | samtools sort - motherChr7Region samtools index motherChr7Region.bam
Call SNPs:
freebayes -f humanChr7Region.fasta --min-alternate-count 10 motherChr7Region.bam >motherChr7Region.vcf
Run igv, use humanChr7Region.fasta as genome, open motherChr7Region.bam and motherChr7Region.vcf. Looking at the aligned reads and the vcf file, answer the following questions in protocol:
- (a) How many variants were found in the vcf file?
- (b) How many variants are heterozygous and how many are homozygous?
- (c) Are all variants single-nucleotide variants or do you also see some insertions/deletions (indels)?
Also export overall view of the whole region from igv to file motherChr7Region.png.
Submit the following files:
- motherChr7Region.png, motherChr7Region.bam, motherChr7Region.vcf
Task B: UCSC browser
- (a) Where is sequence from regionChr7.fasta located in the browser?
- Go to http://genome-euro.ucsc.edu/, From the blue menu, select Tools->Blat
- Check that blat uses Human, hg38 assembly
- Open regionChr7.fasta in a graphical editor (e.g. gedit), sleect all, paste to BLAT window, run BLAT
- In the table of results, the best result should have identiy close to 100% and span close to 7kb
- For this best result, click on link named Browser
- Report which chromosome and which region you get
- (b) Which gene is located in this region?
- Once you are int he browser, press Default tracks button
- Track named GENCODE contains known genes, shown as rectangles (exanos) connected by lines (introns). Short gene names are next to them.
- Report the name of the gene in the region
- (c) In which tissue is this gene most highly expressed? What is the function of this gene?
- When you click on the gene (possibly twice), you get an information page which starts with a summary of the known function of this gene. Copy the first sentence to your protocol.
- Further down on the gene information page you see RNA-Seq Expression Data (colorful boxplots). Find out which tissues have the highest signal.
- (d) Which SNPs are located in this gene? Which trait do they inflence?
- You can see SNPs in the Common SNPs(151) track, but their IDs appear only after switching this track to pack mode. You can click on each SNPs to see more information and to copy their ID to your protocol.
- Information page of the gene (part c) also describes function of various alleles of this gene (see e.g. part POLYMORPHISM).
- You can also find information about individual SNPs by looking for them by their ID in SNPedia (not required in this task)
Task C: Examining larger vcf files
In this task, we will look at motherChr12.vcf and fatherChr12.vcf files and compute various statistics. You can use command-line tools, such as grep, wc, sort, uniq and Perl one-liners (as in #L02), you write small scripts in Perl or Python (as in #L01 and #L04).
- Write all used commands to your protocol
- If you write any scripts, submit them as well.
Questions:
- (a) How many SNPs are in each file?
- This can be found easily by wc, only make sure to exclude lines with comments
- (b) How many heterozygous SNPs are in each file?
- The last column contains 1|1 for homozygous and either 0|1 or 1|0 for heterozygous SNPs
- Character | has special meaning on command line and in grep patterns, make sure to place it in ' ' and possibly escape it with \
- (c) How many SNP positions are shared between the two files?
- The second column of each file lists the position. We want to compute the size of intersection of the set of positions in motherChr12.vcf and fatherChr12.vcf files
- You can e.g. create temporary files containing only positions from the two files and sort them alphabetically. Then you can find the intersection using comm command with options -1 -2. Alternatively, you can store positions as keys in a hash table (dictionary) in a Perl or Python script.
- (d) List the 5 most frequent pairs of reference/alternate allele in motherChr12.vcf and their frequencies. Do they correspond to transitions or transversions?
- Fourth column contains the reference value, fifth column the alternate value. For example, the first SNP in motherChr12.vcf has a pair C,A.
- For each possible pair of nucleotides, find how many times it occurs in the motherChr12.vcf
- For example, pair C,A occurs 6894 times
- Then sort the pairs by their frequencies and report 5 most frequent pairs
- Mutations can be classified as transitions and transversions. Transitions change purine to purine or pyrimidine to pyrimidine, transversion change a purine to pyrimidine or vice versa. For example, pair C,A is a transversion changing pyrimidine C to purine A. Which of these most frequent pairs correspond to transitions and which to transversions?
- To count pairs without writing scripts, you can create a temporary file containing only columns 4 and 5 (without comments), and then use commands sort and uniq to count each pair.
- (e) Which parts of the chromosome have the highest and lowest number of SNPs in motherChr12.vcf?
- First create a list of windows of size 100kb covering the whole chromosome 12 using these two commands:
- perl -le 'print "chr12\t133275309"' > humanChr12.size
- bedtools makewindows -g humanChr12.size -w 100000 -i srcwinnum > humanChr12-windows.bed
- Then count SNPs in each window using this command:
- bedtools coverage -a humanChr12-windows.bed -b motherChr12.vcf > motherChr12-windows.tab
- Find out which column of the resulting file contains the number of SNPs per window, e.g. by reading the documentation obtained by command bedtools coverage -h
- Sort according to the column with SNP number, look at the first and last line of the sorted file
- For checking: the second highest count is 387 in window with coordinates 20,800,000-20,900,000
- First create a list of windows of size 100kb covering the whole chromosome 12 using these two commands:
L08
Program for today: basics of R (applied to biology examples)
- very short intro as a lecture
- tutorial as HW: read a bit of text, try some commands, extend/modify them as requested
In this course we cover several languages popular for scripting in bioinformatics: Perl, Python, R
- their capabilities overlap, many extensions emulate strengths of one in another
- choose a language based on your preference, level of knowledge, existing code for the task, rest of the team
- quickly learn a new language if needed
- also possibly combine, e.g. preprocess data in Perl or Python, then run statistical analyses in R, automate entire pipeline with bash or make
Introduction
- R is an open-source system for statistical computing and data visualization
- Programming language, command-line interface
- Many built-in functions, additional libraries
- For example http://bioconductor.org/ for bioinformatics
- We will concentrate on useful commands rather than language features
Working in R
- Run command R, type commands in command-line interface
- supports history of commands (arrows, up and down, Ctrl-R) and completing command names with tab key
> 1+2 [1] 3
- Write a script to file, run it from command-line: R --vanilla --slave < file.R
- Use rstudio to open a graphics IDE [23]
- Windows with editor of R scripts, console, variables, plots
- Ctrl-Enter in editor executes current command in console
x=c(1:10) plot(x,x*x)
- ? plot displays help for plot command
Suggested workflow
- work interactively in Rstudio or on command line, try various options
- select useful commands, store in a script
- run script automatically on new data/new versions, potentially as a part of a bigger pipeline
Additional information
- Official tutorial
- Seefeld, Linder: Statistics Using R with Biological Examples (pdf book)
- Patrick Burns: The R Inferno (intricacies of the language)
- Other books
Gene expression data
- Gene expression: DNA->mRNA->protein
- Level of gene expression: Extract mRNA from a cell, measure amounts of mRNA
- Technologies: microarray, RNA-seq
Gene expression data
- Rows: genes
- Columns: experiments (e.g. different conditions or different individuals)
- Each value is expression of a gene, i.e. relative amount of mRNA for this gene in the sample
We will use microarray data for yeast:
- Strassburg, Katrin, et al. "Dynamic transcriptional and metabolic responses in yeast adapting to temperature stress." Omics: a journal of integrative biology 14.3 (2010): 249-259. [24]
- Downloaded from GEO database [25]
- Data already preprocessed: normalization, log2, etc
- We have selected only cold conditions, genes with absolute change at least 1
- Data: 2738 genes, 8 experiments in a time series, yeast moved from normal temperature 28 degrees C to cold conditions 10 degrees C, samples taken after 0min, 15min, 30min, 1h, 2h, 4h, 8h, 24h in cold
HW08
Submitting
In this homework, try to read text, execute given commands, potentially trying some small modifications.
- Then do tasks A-D, submit required files (3x .png)
- In your protocol, enter commands used in tasks A-D, with explanatory comments in more complicated situations
- In task B also enter required output to protocol
- Protocol template in /tasks/hw08/protocol.txt
First steps
- Type a command, R writes the answer, e.g.:
> 1+2 [1] 3
- We can store values in variables and use them later on
> # The size of the sequenced portion of cow's genome, in millions of base pairs > Cow_genome_size <- 2290 > Cow_genome_size [1] 2290 > Cow_chromosome_pairs <- 30 > Cow_avg_chrom <- Cow_genome_size / Cow_chromosome_pairs > Cow_avg_chrom [1] 76.33333
Surprises:
- dots are used as parts of id's, e.g. read.table is name of a single function (not method for object read)
- assignment via <- or =
- careful: a<-3 is an assignment, a < -3 is a comparison
- vectors etc are indexed from 1, not from 0
Vectors, basic plots
- Vector is a sequence of values of the same type (all are numbers or all are strings or all are booleans)
# Vector can be created from a list of numbers by function named c
a <- c(1,2,4)
a
# prints [1] 1 2 4
# c also concatenates vectors
c(a,a)
# prints [1] 1 2 4 1 2 4
# Vector of two strings
b <- c("hello", "world")
# Create a vector of numbers 1..10
x <- 1:10
x
# prints [1] 1 2 3 4 5 6 7 8 9 10
Vector arithmetics
- Operations applied to each member of the vector
x <- 1:10 # Square each number in vector x x*x # prints [1] 1 4 9 16 25 36 49 64 81 100 # New vector y: logarithm of a number in x squared y <- log(x*x) y # prints [1] 0.000000 1.386294 2.197225 2.772589 3.218876 3.583519 3.891820 4.158883 # [9] 4.394449 4.605170 # Draw graph of function log(x*x) for x=1..10 plot(x,y) # The same graph but use lines instead of dots plot(x,y,type="l") # Addressing elements of a vector: positions start at 1 # Second element of the vector y[2] # prints [1] 1.386294 # Which elements of the vector satisfy certain condition? (vector of logical values) y>3 # prints [1] FALSE FALSE FALSE FALSE TRUE TRUE TRUE TRUE TRUE TRUE # write only those elements from y that satisfy the condition y[y>3] # prints [1] 3.218876 3.583519 3.891820 4.158883 4.394449 4.605170 # we can also write values of x such that values of y satisfy the condition... x[y>3] # prints [1] 5 6 7 8 9 10
- Alternative plotting facilities: ggplot2 library, lattice library
Task A
- Create a plot of the binary logarithm with dots in the graph more densely spaced (from 0.1 to 10 with step 0.1)
- Store it in file log.png and submit this file
- Hints:
- Create x and y by vector arithmetics
- To compute binary logarithm check help ? log
- Before running plot, use command png("log.png") to store the result, afterwards call dev.off() to close the file (in rstudio you can also export plots manually)
Data frames and simple statistics
- Data frame: a table similar to spreadsheet, each column is a vector, all are of the same length
- We will use a table with the following columns:
- The size of a genome, in millions of nucleotides
- Number of chromosome pairs
- GC content
- Taxonomic group mammal or fish
- Stored in CSV format, columns separated by tabs.
- Data: Han et al Genome Biology 2008 [26]
Species Size Chrom GC Group Human 2850 23 40.9 mammal Chimpanzee 2750 24 40.7 mammal Macaque 2650 21 40.7 mammal Mouse 2480 20 41.7 mammal ... Tetraodon 187 21 45.9 fish ...
# reading a frame from file
a<-read.table("/tasks/hw08/genomes.csv", header = TRUE, sep = "\t");
# column with name size
a$Size
# Average chromosome length: divide size by the number of chromosomes
a$Size/a$Chrom
# Add average chromosome length as a new column to frame a
a<-cbind(a,AvgChrom=a$Size/a$Chrom)
# Scatter plot of average chromosome length vs GC content
plot(a$AvgChrom, a$GC)
# Compactly display structure of a
# (good for checking that import worked etc)
str(a)
# display mean, median, etc. of each column
summary(a);
# average genome size
mean(a$Size)
# average genome size for mammals
mean(a$Size[a$Group=="mammal"])
# Standard deviation
sd(a$Size)
# Histogram of genome sizes
hist(a$Size)
Task B
- Divide frame a to two frames, one for mammals, one for fish. Hint:
- Try command a[c(1,2,3),]. What is it doing?
- Try command a$Group=="mammal".
- Combine these two commands to get rows for all mammals and store the frame in a new variable, then repeat for fish
- Use a general approach which does not depend on the exact number and ordering of rows in the table.
- Run the command summary separately for mammals and for fish. Which of their characteristics are different?
- Write output and your conclusion to the protocol
Task C
- Draw a graph comparing genome size vs GC content; use different colors for points representing mammals and fish
- Submit the plot in file genomes.png
- To draw the graph, you can use one of the options below, or find yet another way
- Option 1: first draw mammals with one color, then add fish in another color
- Color of points can be changed by: plot(1:10,1:10, col="red")
- After plot command you can add more points to the same graph by command points, which can be used similarly as plot
- Warning: command points does not change the ranges of x and y axes. You have to set these manually so that points from both groups are visible. You can do this using options xlim and ylim, e.g. plot(x,y, col="red", xlim=c(1,100), ylim=c(1,100))
- Option 2: plot both mammals and fish in one plot command, and give it a vector of colors, one for each point
- plot(1:10,1:10,col=c(rep("red",5),rep("blue",5))) will plot the first 5 points red and the last 5 points blue
- Bonus task: add a legend to the plot, showing which color is mammal and which is fish
Expression data and clustering
Data here is bigger, better to use plain R rather than rstudio (limited server CPU/memory)
# Read gene expression data table
a <- read.table("/tasks/hw08/microarray.csv", header = TRUE, sep = "\t", row.names=1)
# Visual check of the first row
a[1,]
# plot starting point vs. situation after 1 hour
plot(a$cold_0min,a$cold_1h)
# to better see density in dense clouds of points, use this plot
smoothScatter(a$cold_15min, a$cold_1h)
# outliers away from diagonal in the plot above are most strongly differentially expressed genes
# these are more easy to see in MA plot:
# x-axis: average expression in the two conditions
# y-axis: difference between values (they are log-scale, so difference 1 means 2-fold)
smoothScatter((a$cold_15min+a$cold_1h)/2, a$cold_15min-a$cold_1h)
Clustering is a wide group of methods that split data points into groups with similar properties
- We will group together genes that have a similar reaction to cold, i.e. their rows in gene expression data matrix have similar values
We will consider two simple clustering methods
- K means clustering splits points (genes) into k clusters, where k is a parameter given by the user. It finds a center of each cluster and tries to minimize the sum of distances from individual points to the center of their cluster. Note that this algorithm is randomized so you will get different clusters each time.
- Hierarchical clustering puts all data points (genes) to a hierarchy so that smallest subtrees of the hierarchy are the most closely related groups of points and these are connected to bigger and more loosely related groups.
# Heatmap: creates hierarchical clustering of rows # then shows every value in the table using color ranging from red (lowest) to white (highest) # Computation may take some time heatmap(as.matrix(a),Colv=NA) # Previous heatmap normalized each row, the next one uses data as they are: heatmap(as.matrix(a),Colv=NA,scale="none")
# k means clustering to 7 clusters k = 7 cl <- kmeans(a,k) # each gene has assigned a cluster (number between 1 and k) cl$cluster # draw only cluster number 3 out of k heatmap(as.matrix(a[cl$cluster==3,]),Rowv=NA, Colv=NA) # reorder genes in the table according to cluster heatmap(as.matrix(a[order(cl$cluster),]),Rowv=NA, Colv=NA) # compare overall column means with column means in cluster 3 # function apply uses mean on every column (or row if 2 changed to 1) apply(a,2,mean) # now means within cluster apply(a[cl$cluster==3,],2,mean) # clusters have centers which are also computed as means # so this is the same as previous command cl$centers[3,]
Task D
- Draw a plot in which x-axis is time and y-axis is the expression level and the center of each cluster is shown as a line
- use command matplot(x,y,type="l") which gets two matrices x and y and plots columns of x vs columns of y
- matplot(,y,type="l") will use numbers 1,2,3... as columns of the missing matrix x
- create y from cl$centers by applying function t (transpose)
- to create an appropriate matrix x, create a vector of times for individual experiments in minutes or hours (do it manually, no need to parse column names automatically)
- using functions rep and matrix you can create a matrix x in which this vector is used as every column
- then run matplot(x,y,type="l")
- since time points are not evenly spaced, it would be better to use logscale: matplot(x,y,type="l",log="x")
- to avoid log(0), change the first timepoint from 0min to 1min
- Submit file clusters.png with your final plot
L09
Topic of this lecture are statistical tests in R.
- Beginners in statistics: listen to lecture, then do tasks A, B, C
- If you know basics of statistical tests, do tasks B, C, D
- More information on this topic in 1-EFM-340 Počítačová štatistika
Introduction to statistical tests: sign test
- [27]
- Two friends A and B have played their favourite game n=10 times, A has won 6 times and B has won 4 times.
- A claims that he is a better player, B claims that such a result could easily happen by chance if they were equally good players.
- Hypothesis of player B is called null hypothesis that the pattern we see (A won more often than B) is simply a result of chance
- Null hypothesis reformulated: we toss coin n times and compute value X: the number of times we see head. The tosses are independent and each toss has equal probability of being head or tail
- Similar situation: comparing programs A and B on several inputs, counting how many times is program A better than B.
# simulation in R: generate 10 psedorandom bits
# (1=player A won)
sample(c(0,1), 10, replace = TRUE)
# result e.g. 0 0 0 0 1 0 1 1 0 0
# directly compute random variable X, i.e. sum of bits
sum(sample(c(0,1), 10, replace = TRUE))
# result e.g. 5
# we define a function which will m times repeat
# the coin tossing experiment with n tosses
# and returns a vector with m values of random variable X
experiment <- function(m, n) {
x = rep(0, m) # create vector with m zeroes
for(i in 1:m) { # for loop through m experiments
x[i] = sum(sample(c(0,1), n, replace = TRUE))
}
return(x) # return array of values
}
# call the function for m=20 experiments, each with n tosses
experiment(20,10)
# result e.g. 4 5 3 6 2 3 5 5 3 4 5 5 6 6 6 5 6 6 6 4
# draw histograms for 20 experiments and 1000 experiments
png("hist10.png") # open png file
par(mfrow=c(2,1)) # matrix of plots with 2 rows and 1 column
hist(experiment(20,10))
hist(experiment(1000,10))
dev.off() # finish writing to file
- It is easy to realize that we get binomial distribution (binomické rozdelenie)
- P-value of the test is the probability that simply by chance we would get k the same or more extreme than in our data.
- In other words, what is the probability that in 10 tosses we see head 6 times or more (one sided test)
- If the p-value is very small, say smaller than 0.01, we reject the null hypothesis and assume that player A is in fact better than B
# computing the probability that we get exactly 6 heads in 10 tosses
dbinom(6, 10, 0.5) # result 0.2050781
# we get the same as our formula above:
7*8*9*10/(2*3*4*(2^10)) # result 0.2050781
# entire probability distribution: probabilities 0..10 heads in 10 tosses
dbinom(0:10, 10, 0.5)
# [1] 0.0009765625 0.0097656250 0.0439453125 0.1171875000 0.2050781250
# [6] 0.2460937500 0.2050781250 0.1171875000 0.0439453125 0.0097656250
# [11] 0.0009765625
#we can also plot the distribution
plot(0:10, dbinom(0:10, 10, 0.5))
barplot(dbinom(0:10,10,0.5))
#our p-value is sum for 6,7,8,9,10
sum(dbinom(6:10,10,0.5))
# result: 0.3769531
# so results this "extreme" are not rare by chance,
# they happen in about 38% of cases
# R can compute the sum for us using pbinom
# this considers all values greater than 5
pbinom(5, 10, 0.5, lower.tail=FALSE)
# result again 0.3769531
# if probability is too small, use log of it
pbinom(9999, 10000, 0.5, lower.tail=FALSE, log.p = TRUE)
# [1] -6931.472
# the probability of getting 10000x head is exp(-6931.472) = 2^{-100000}
# generating numbers from binomial distribution
# - similarly to our function experiment
rbinom(20, 10, 0.5)
# [1] 4 4 8 2 6 6 3 5 5 5 5 6 6 2 7 6 4 6 6 5
# running the test
binom.test(6, 10, p = 0.5, alternative="greater")
#
# Exact binomial test
#
# data: 6 and 10
# number of successes = 6, number of trials = 10, p-value = 0.377
# alternative hypothesis: true probability of success is greater than 0.5
# 95 percent confidence interval:
# 0.3035372 1.0000000
# sample estimates:
# probability of success
# 0.6
# to only get p-value run
binom.test(6, 10, p = 0.5, alternative="greater")$p.value
# result 0.3769531
Comparing two sets of values: Welch's t-test
- Let us now consider two sets of values drawn from two normal distributions with unknown means and variances
- The null hypothesis of the Welch's t-test is that the two distributions have equal means
- The test computes test statistics (in R for vectors x1, x2):
- (mean(x1)-mean(x2))/sqrt(var(x1)/length(x1)+var(x2)/length(x2))
- This test statistics is approximately distributed according to Student's t-distribution with the degree of freedom obtained by
n1=length(x1) n2=length(x2) (var(x1)/n1+var(x2)/n2)**2/(var(x1)**2/((n1-1)*n1*n1)+var(x2)**2/((n2-1)*n2*n2))
- Luckily R will compute the test for us simply by calling t.test
x1 = rnorm(6, 2, 1) # 2.70110750 3.45304366 -0.02696629 2.86020145 2.37496993 2.27073550 x2 = rnorm(4, 3, 0.5) # 3.258643 3.731206 2.868478 2.239788 > t.test(x1,x2) # t = -1.2898, df = 7.774, p-value = 0.2341 # alternative hypothesis: true difference in means is not equal to 0 # means 2.272182 3.024529 x2 = rnorm(4, 5, 0.5) # 4.882395 4.423485 4.646700 4.515626 t.test(x1,x2) # t = -4.684, df = 5.405, p-value = 0.004435 # means 2.272182 4.617051 # to get only p-value, run t.test(x1,x2)$p.value
We will apply Welch's t-test to microarray data
- Data from GEO database [28], publication [29]
- Abbott et al 2007: Generic and specific transcriptional responses to different weak organic acids in anaerobic chemostat cultures of Saccharomyces cerevisiae
- gene expression measurements under 5 conditions:
- reference: yeast grown in normal environment
- 4 different acids added so that cells grow 50% slower (acetic, propionic, sorbic, benzoic)
- from each condition (reference and each acid) we have 3 replicates
- together our table has 15 columns (3 replicates from 5 conditions)
- 6398 rows (genes)
- We will test statistical difference between the reference condition and one of the acids (3 numbers vs other 3 numbers)
- See Task B in #HW09
Multiple testing correction
- When we run t-tests on the reference vs. acetic acid on all 6398 genes, we get 118 genes with p-value<=0.01
- Purely by chance this would happen in 1% of cases (from definition of p-value)
- So purely by chance we would expect to get about 64 genes with p-value<=0.01
- So perhaps roughly half of our detected genes (maybe less, maybe more) are false positives
- Sometimes false positives may even overwhelm results
- Multiple testing correction tries to limit the number of false positives among results of multiple statistical tests
- Many different methods
- The simplest one is Bonferroni correction, where the threshold on p-value is divided by the number of tested genes, so instead of 0.01 we use 0.01/6398 = 1.56e-6
- This way the expected overall number of false positives in the whole set is 0.01 and so the probability of getting even a single false positive is also at most 0.01 (by Markov inequality)
- We could instead multiply all p-values by the number of tests and apply the original threshold 0.01 - such artificially modified p-values are called corrected
- After Bonferroni correction we get only 1 significant gene
# the results of p-tests are in vector pa of length 6398 # manually multiply p-values by length(pa), count those that have value <=0.01 sum(pa * length(pa) < 0.01) # in R you can use p.adjust form multiple testing correction pa.adjusted = p.adjust(pa, method ="bonferroni") # this is equivalent to multiplying by the length and using 1 if the result > 1 pa.adjusted = pmin(pa*length(pa),rep(1,length(pa))) # there are less conservative multiple testing correction methods, e.g. Holm's method # but in this case we get almost the same results pa.adjusted2 = p.adjust(pa, method ="holm")
- Other frequently used correction is false discovery rate (FDR), which is less strict and controls the overall proportion of false positives among results
HW09
- Do either tasks A,B,C (beginners) or B,C,D (more advanced). If you really want, you can do all four for bonus credit.
- In your protocol write used R commands with brief comments on your approach.
- Submit required plots with filenames as specified.
- For each task also include results as required and a short discussion commenting the results/plots you have obtained. Is the value of interest increasing or decreasing with some parameter? Are the results as expected or surprising?
- Outline of protocol is in /tasks/hw09/protocol.txt
Task A: sign test
- Consider a situation in which players played n games, out of which a fraction of q were won by A (example in lecture corresponds to q=0.6 and n=10)
- Compute a table of p-values for n=10,20,...,90,100 and for q=0.6, 0.7, 0.8, 0.9
- Plot the table using matplot (n is x-axis, one line for each value of q)
- Submit the plot in sign.png
- Discuss the values you have seen in the plot / table
Outline of the code:
# create vector rows with values 10,20,...,100
rows=(1:10)*10
# create vector columns with required values of q
columns=c(0.6, 0.7, 0.8, 0.9)
# create empty matrix of pvalues
pvalues = matrix(0,length(rows),length(columns))
# TODO: fill in matrix pvalues using binom.test
# set names of rows and columns
rownames(pvalues)=rows
colnames(pvalues)=columns
# careful: pvalues[10,] is now 10th row, i.e. value for n=100,
# pvalues["10",] is the first row, i.e. value for n=10
# check that for n=10 and q=0.6 you get p-value 0.3769531
pvalues["10","0.6"]
# create x-axis matrix (as in HW08, part D)
x=matrix(rep(rows,length(columns)),nrow=length(rows))
# matplot command
png("sign.png")
matplot(x,pvalues,type="l",col=c(1:length(columns)),lty=1)
legend("topright",legend=columns,col=c(1:length(columns)),lty=1)
dev.off()
Task B: Welch's t-test on microarray data
- Read table with microarray data, transform it to log scale, then work with table a:
input=read.table("/tasks/hw09/acids.tsv", header=TRUE, row.names=1)
a = log(input)
- Columns 1,2,3 are reference, columns 4,5,6 acetic acid, 7,8,9 benzoate, 10,11,12 propionate, and 13,14,15 sorbate
- Write a function my.test which will take as arguments table a and 2 lists of columns (e.g. 1:3 and 4:6) and will run for each row of the table Welch's t-test of the first set of columns vs the second set. It will return the resulting vector of p-values
- For example by calling pa <- my.test(a, 1:3, 4:6) we will compute p-values for differences between reference and acetic acid (computation may take some time)
- The first 5 values of pa should be
> pa[1:5] [1] 0.94898907 0.07179619 0.24797684 0.48204100 0.23177496
- Run the test for all four acids
- Report how many genes were significant with p-value <= 0.01 for each acid
- See Vector arithmetics in HW08
- You can count TRUE items in a vector of booleans by sum, e.g. sum(TRUE,FALSE,TRUE) is 2
- Report how many genes are significant for both acetic and benzoate acids? (logical and is written as &)
Task C: multiple testing correction
Run the following snippet of code, which works on the vector of p-values pa obtained for acetate in task B
# adjusts vectors of p-vales from tasks B for using Bonferroni correction pa.adjusted = p.adjust(pa, method ="bonferroni") # add this adjusted vector to frame a a <- cbind(a, pa.adjusted) # create permutation ordered by pa.adjusted oa = order(pa.adjusted) # select from table five rows with the lowest pa.adjusted (using vector oa) # and display columns containing reference, acetate and adjusted p-value a[oa[1:5],c(1:6,16)]
You should get output like this:
ref1 ref2 ref3 acetate1 acetate2 acetate3 pa.adjusted SUL1 7.581312 7.394985 7.412040 2.1633230 2.05412373 1.9169226 0.004793318 YMR244W 2.985682 2.975530 3.054001 0.3364722 0.33647224 0.1823216 0.188582576 DIP5 6.943991 7.147795 7.296955 0.6931472 0.09531018 0.5306283 0.253995075 YLR460C 5.620401 5.801212 5.502482 3.2425924 3.48431229 3.3843903 0.307639012 HXT4 2.821379 3.049273 2.772589 7.7893717 8.24446541 8.3041980 0.573813502
Do the same procedure for benzoate p-values and report the result (in your table, report both p-values and expression levels for bezoate, not acetate). Comment the results for both acids.
Task D: volcano plot, test on data generated from null hypothesis
Draw a volcano plot for the acetate data
- x-axis of this plot is the difference in the mean of reference and mean of acetate.
- You can compute row means of a matrix by rowMeans.
- y-axis is -log10 of the p-value (use original p-values before multiple testing correction)
- You can quickly see genes which have low p-values (high on y-axis) and also big difference in mean expression between the two conditions (far from 0 on x-axis). You can also see if acetate increases or decreases expression of these genes.
Now create a simulated dataset sharing some features of the real data but observing the null hypothesis that the mean of reference and acetate are the same for each gene
- Compute vector m of means for columns 1:6 from matrix a
- Compute vectors sr and sa of standard deviations for reference columns and for acetate columns respectively
- You can compute standard deviation for each row of a matrix by apply(some.matrix, 1, sd)
- For each i in 1:6398, create three samples from normal distribution with mean m[i] and standard deviation sr[i], and three samples with mean m[i] and deviation sa[i]
- Use function rnorm
- On the resulting matrix apply Welch's t-test and draw the volcano plot.
- How many random genes have p-value <=0.01? Is it roughly what we would expect under the null hypothesis?
Draw histogram of p-values from the real data (reference vs acetate) and from random data. Describe how they differ. Is it what you would expect?
- use function hist
Submit plots volcano-real.png, volcano-random.png, hist-real.png, hist-random.png (real for real expression data and random for generated data)
L10
Today we will work with AWS. Please use credentials which were sent to you via email and follows steps in here (there is a cursor in each screen): https://docs.google.com/presentation/d/1GBDErp5xhrV2zLF5kKdwnOAjtmDEFN0pw3RFval419s/edit#slide=id.p
The goal is to have '.aws/credentials file in your homefolder. Sometimes these credentials expire, follow same steps to refresh them (copy them again).
There is aws command line tools installed on vyuka machine (on your machine use pip install awscli).
Now you need to do two things:
Check if you can download a file from our S3 datastore (think of S3 as some remote file storage).
aws s3 ls s3://idzbucket2 - should give you a big list of files aws s3 cp s3://idzbucket2/splitaa splitaa - should download a file to your machine aws s3 cp s3://idzbucket2/splitaa - - will print file in your console (no need to do this).
Check if you can create a bucket (your storage area, pick your own name, must be globally unique): aws s3 mb s3://mysuperawesomebucket
We will be using MapReduce in this session. (It is kind of outdated concept, but simple enough for us and runs out of box on AWS. If you ever want to use BigData in practice, try something more modern like Apache Beam. And avoid PySpark if you can.)
For tutorial on mapreduce check out [30] or [31].
You are given basic template with comments at tasks/example_job.py and you can run it locally as python3 example_job.py <input file> -o <output_dir> or in cloud as python3 example_job.py -r emr s3://idzbucket2 --num-core-instances 4 -o s3://<your bucket>/<some directory> (but for testing we recommend smaller sample python3 example_job.py -r emr s3://idzbucket2/splita* --num-core-instances 4 -o s3://<your bucket>/<some directory>).
You can download output using aws s3 ls s3://<your bucket>/<some directory>/ and aws s3 cp s3://<your bucket>/<some directory>/ . --recursive
If you want to watch progress: Click on AWS Console button workbench (vocareum). Set region (top right) to Oregon. Click on services, then EMR. Click on the job, which is running, then Steps, view logs, syslog.
HW10
Task 1:
Count number of occurences of each 4-mer in provided data.
Task 2:
Count number of pairs of reads which overlaps in exactly 30 bases (end of one read overlaps beginning of second read). You can ignore reverse complement. Hints:
- Try counting pairs for each 30-mer first.
- You can yield something structured from Mapper (e.g. tuple).
- There is a two step mapreduce, which can help you with final summation: https://pythonhosted.org/mrjob/guides/writing-mrjobs.html
For both tasks, submit source code and the result, when run on whole dataset (s3://idzbucket2).
Code is expected to use MRJob framework presented in lecture.